Overview
Panorama is a freely available, open-source web server database application for targeted proteomics assays that integrates into a Skyline proteomics workflow. It has been implemented as a module within
LabKey Server, an open-source bioinformatics data management platform with extensive support for proteomics data and a security model rich enough to support clinical studies.
Panorama can be installed by laboratories and organizations on their own servers. You can visit the LabKey Server installation documentation:
https://www.labkey.org/wiki/home/Documentation/page.view?name=installServerDemo for the available installation options. For this tutorial you will use the Panorama server hosted at the University of Washington
panoramaweb.org where users can request free projects and have full administrative rights to configure data organization and security.
This tutorial is an introduction to Panorama and covers the following areas:
- Requesting a project on panoramaweb.org
- Data organization and folder management in Panorama
- Publishing Skyline documents to Panorama
- Data display options and searching results uploaded to Panorama
- Providing access to collaborators and other groups
Getting Started
- To start this tutorial, download the following ZIP file:
- Extract the files in it to a folder on your computer, such as
C:\Users\userName\Documents
- This will create a new folder:
C:\Users\userName\Documents\PanoramaSharing
It will contain all the files necessary for this tutorial.
Requesting a Project on panoramaweb.org
Open the Study9S_Site52_v3.sky file in Skyline either by double-clicking on the file in Windows Explorer or by going through the
File > Open menu in Skyline. This file contains results that are part of a recent CPTAC multi-site SRM-MS system suitability study (Abbatiello et al., Mol. Cell. Proteomics 2013). The data in this document was collected at Site 52. To publish this document to Panorama, click on the
 Upload to Panorama
Upload to Panorama button in the toolbar, as circled below. Alternatively, on the
File menu, click
Upload to Panorama.
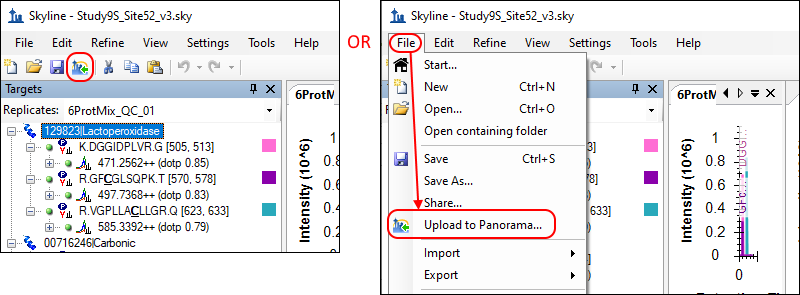
Since you have not yet registered a Panorama server in Skyline, you will see the following message:
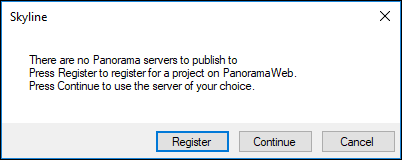
- If you have an existing account on a Panorama server (or LabKey Server), and you would like to use that, click Continue and enter the server details in the Edit Server form Skyline presents. Skip to the next section of this tutorial.
- If you do not have an existing Panorama account, click Register to request a project on the PanoramaWeb server hosted at the University of Washington. This will open the project sign-up page on panoramaweb.org in a web-browser as shown below.
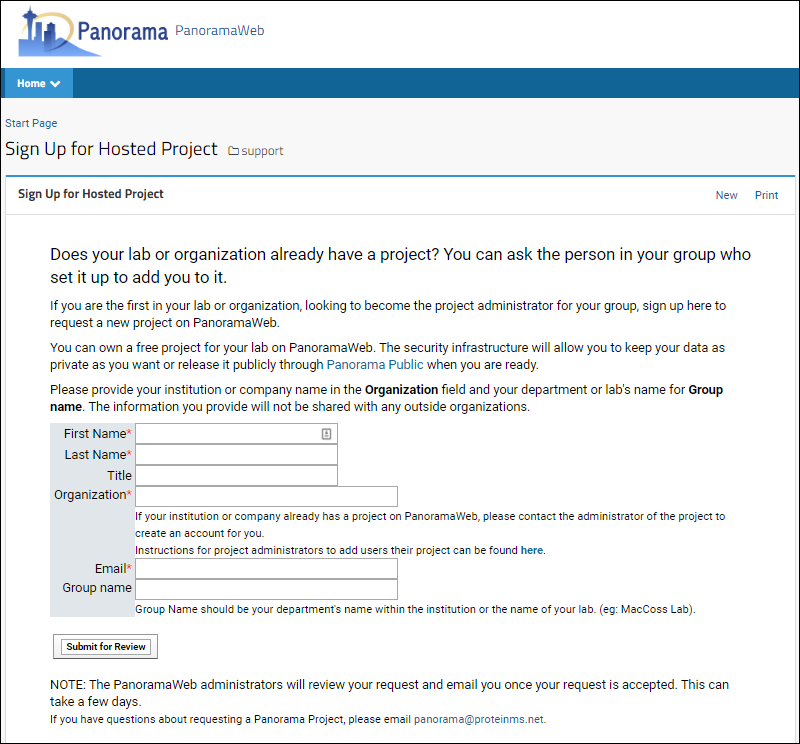
- Fill out the form and be sure to enter a Group name.
- Click Submit for Review.
The group name is used to create projects on panoramaweb.org. The Panorama team reviews all project requests and, upon approval, creates a project with the requested group name. If you are looking for a space to work through this tutorial and explore Panorama, please enter "Tutorials" in the Group name, and a folder with full administrative rights will be created for you in the
Tutorials project on panoramaweb.org. Otherwise, enter your preferred project name. Once your project or folder has been created, you will receive a welcoming email with a link to setup a password for your account. After you have completed the registration process you can sign-in to panoramaweb.org. On the home page, click the
Projects tab to see a list of all the projects to which you have access on panoramaweb.org. If you entered "Tutorials" in the project sign-up form, you will see the
Tutorials project where a folder was just created for you.
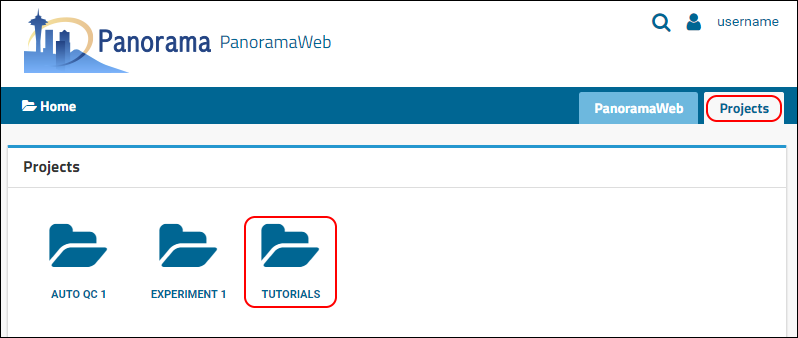
Click on the
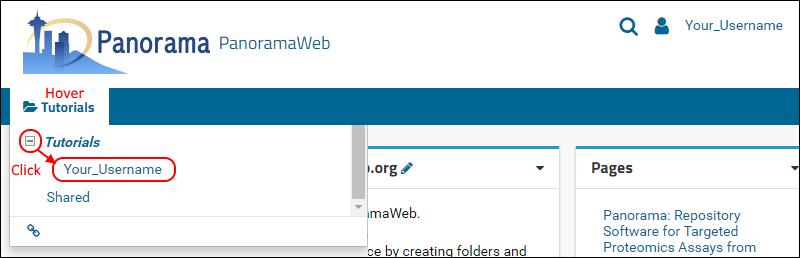
Tutorials project to go to the project homepage. On the project homepage, hover over the project name in the menu bar below the Panorama icon, click the
to open the Tutorials node, then click on the folder that was created for you (e.g. ‘Your_Username’ in the following screenshot). You will have full administrative rights in your folder, so you will be able to create a folder structure for organizing your Skyline documents, and configure user access to the sub-folders.
Creating a Subfolder in Panorama
Skyline documents cannot be uploaded to the root project folder on panoramaweb.org or your folder in the ‘Tutorials’ project. You will create a sub-folder for uploading the Skyline document that you opened earlier. As a project administrator, you would typically organize your project workspace to have subfolders for each project and/or each researcher in your laboratory or organization. Follow these steps to create a "system suitability" sub-folder in your folder in the “Tutorials” project:
- Navigate to "Your_Username" folder home page, if you are not already there.
- Hover over the project name in the menu bar below the Panorama icon and click on the new folder icon shown in the image below.

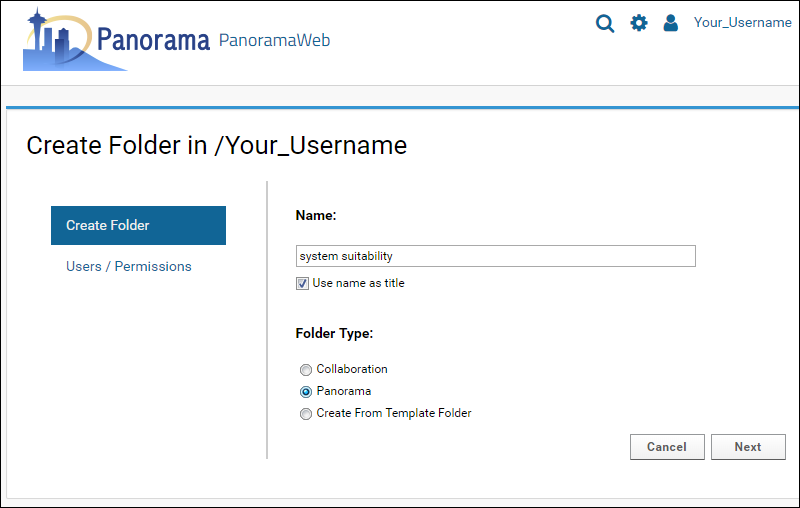
- In the Create Folder form, enter "system suitability" in the folder Name field.
- Select the Panorama option under Folder Type. This is the folder type that should be selected for all workflows supported by Skyline (SRM-MS, MS1 filtering or MS2 based projects).
- Click Next.
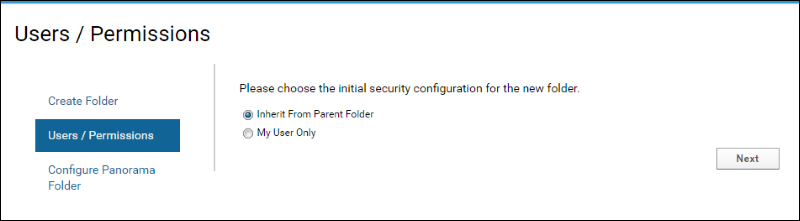
- On the "Users / Permissions" page accept the default selection and click Next. You will configure user permissions on this folder later.
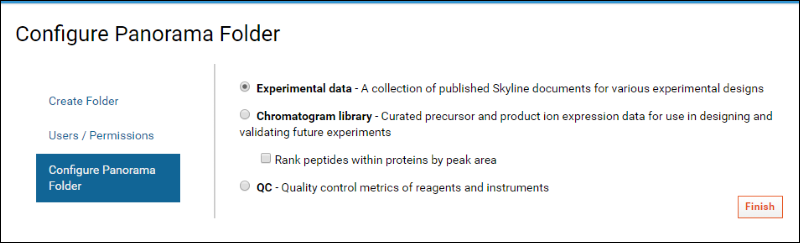
- The next page Configure Panorama Folder asks you to choose the type of Targeted MS folder you would like to create. Panorama offers three choices here. For this tutorial accept the default Experimental data option. This option is meant for folders that serve as a repository of Skyline documents, useful for collaborating, sharing and searching across multiple experiments.
- Click Finish.
You will be taken to the home page of the "system suitability" sub-folder that should look as follows:
You are now ready to upload Skyline documents to this folder.
Publishing Data from Skyline to Panorama
To publish your first Skyline document to PanoramaWeb, perform the following steps:
- Open "Study9S_Site52_v3.sky" in Skyline if it is not already open.
- If the Edit Server form is not already open, open it by clicking the
 Upload to Panorama button, or selecting File > Upload to Panorama and clicking Continue in the popup.
Upload to Panorama button, or selecting File > Upload to Panorama and clicking Continue in the popup.
- In the Edit Server form, enter the email address and password for your account.
- If you are using PanoramaWeb, the server URL should already be entered as https://panoramaweb.org.
- If not, enter the URL to your Panorama/LabKey Server.
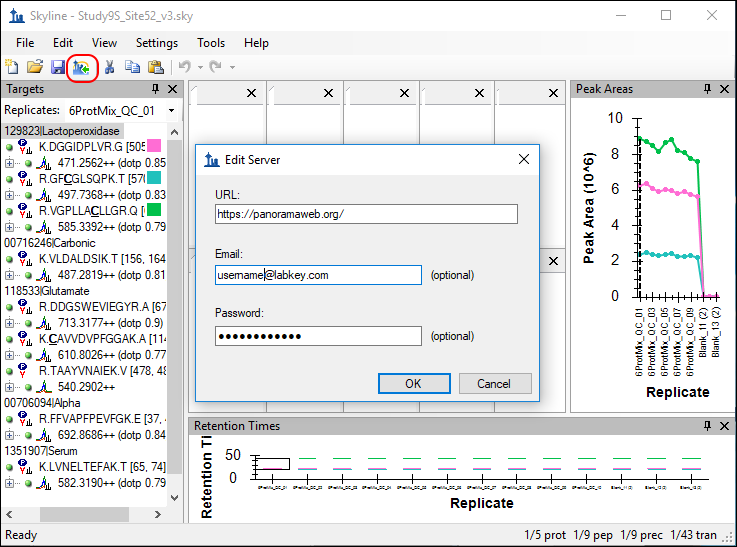
Skyline will now display a form with the folder hierarchy on panoramaweb.org. Only the folders to which you have access will be displayed.
- Navigate to and select the "system suitability" folder.
- Click OK.
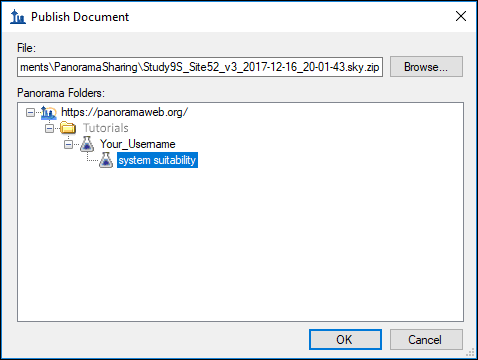
Skyline will create a ZIP archive of the files for your document and upload the ZIP file to panoramaweb.org, where it will be imported into the Panorama database.
A popup message reading "Publish succeeded, would you like to view the file in Panorama?" will appear. Click
Yes to view the document details.
Viewing Data in Panorama
If you are not already viewing the document details, return to your "system suitability" folder, refresh the browser window if necessary, and click the name of the file you just uploaded in the
Targeted MS Runs box.
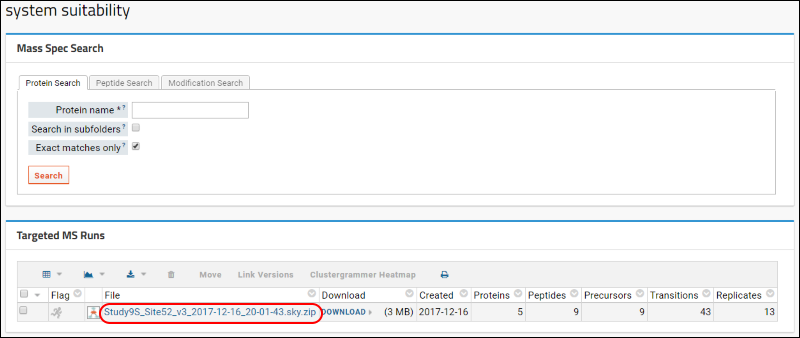
The
Document Summary box shown in image below gives you a summary of the number of proteins, peptides, and transitions in your document.
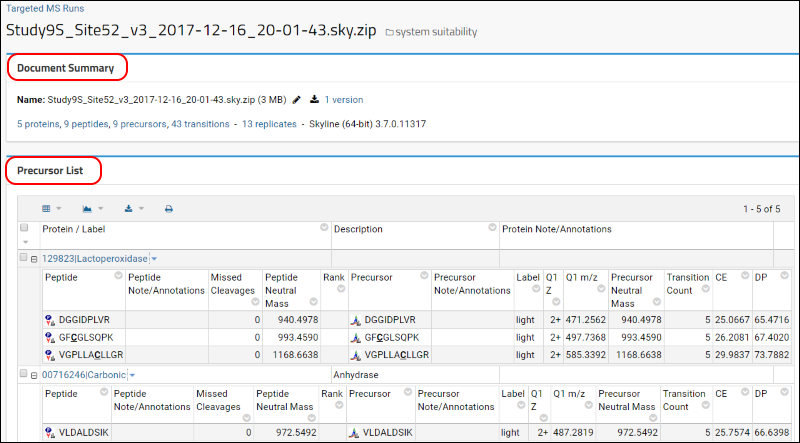
Below the
Document Summary box you can view a list of all the proteins, peptides and precursors in your document, along with some of the information imported from the Skyline document, such as a modified peptide sequence, m/z and charge. You can click on a protein or peptide to view more details, including chromatograms and some of the other graphs that are also available in Skyline. Scroll down on the page and click on the peptide
CAVVDVPFGGAK. This will take you to a page that has the following details for this peptide:
1. Chromatograms to show quantitative data quality
This peptide was measured in 10 QC system suitability replicates and there were 3 blank acquisitions. On this page you can view the chromatograms for this peptide in all the replicates. The image below shows the precursor and transition chromatograms in the replicates QC_01 and QC_02. There is one row of chromatograms for each replicate in your Skyline document, where the first column has the total precursor chromatogram and the subsequent columns have transition chromatograms. If this precursor was measured in a light as well as a heavy-labeled from, transition chromatograms for both the forms would be displayed side-by-side.
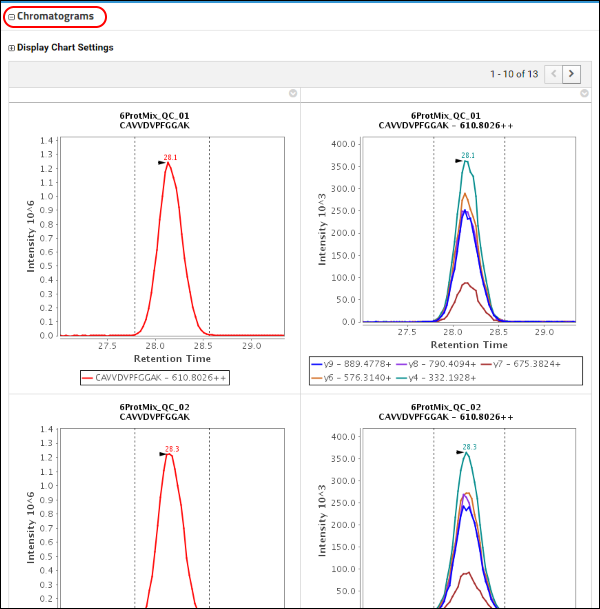
2. Peak Area Replicate Graph
Scroll further down on the page or collapse the Chromatograms box by clicking on the
icon in the header of the box as circled in the image above.
In the
Summary Charts panel, you will now see a graph displaying the peak area for this peptide in the 13 replicates. This graph is similar to what you would see in Skyline for this peptide.
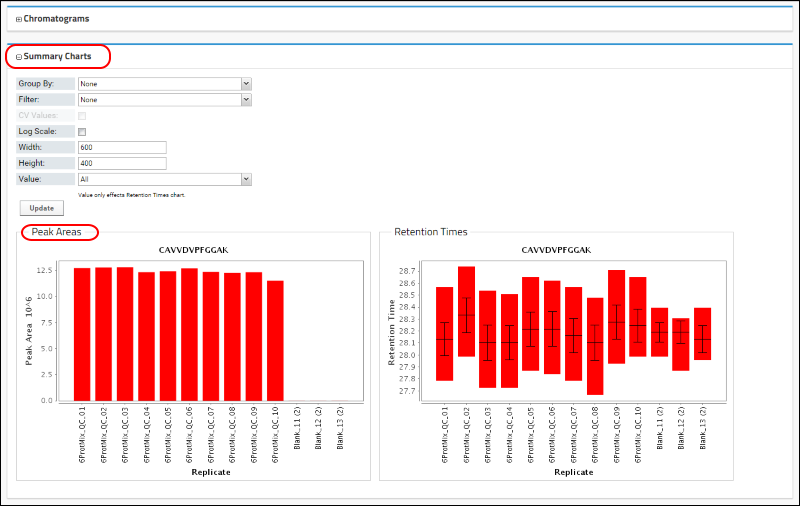
As in Skyline, Panorama also allows grouping peak areas by custom replicate annotations. In this Skyline document researchers used a custom "system suitability" annotation that was set to "true" for the system suitability replicates and "false" for the blank acquisitions. You can group the peak areas by selecting "system suitability" from the pull-down combo-box next to
Group By.
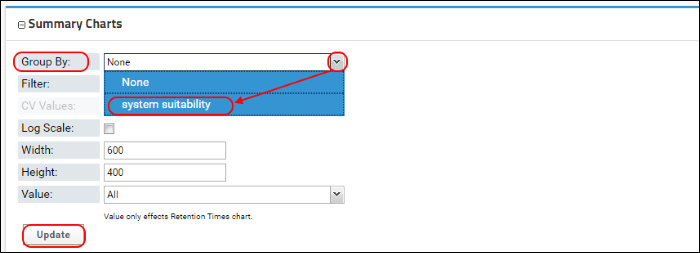
Click
Update to see peak areas grouped by the "system suitability" annotation.
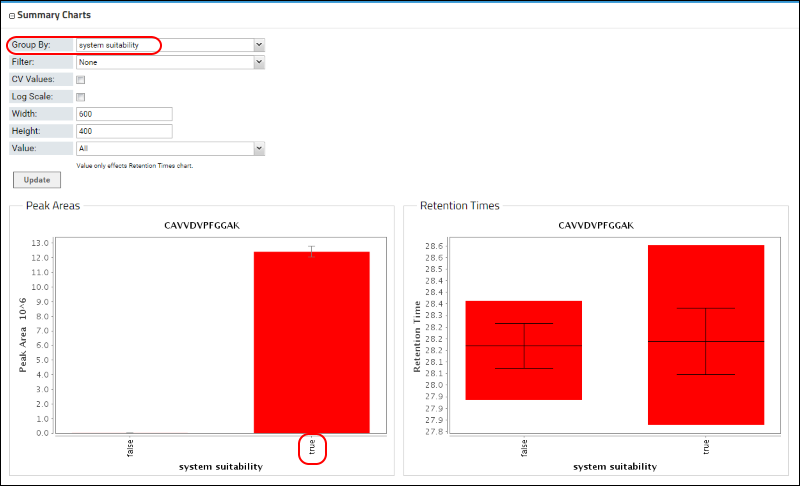
In the graph above, the red bar labeled ‘true’ represents the 10 system suitability runs, and the small error bar indicates that there was very little variability in the measurements for this peptide in the 10 replicates. You can also switch to a Coefficient of Variation (CV) view by checking the
CV Values box and clicking
Update again.
3. Library MS/MS spectrum viewer
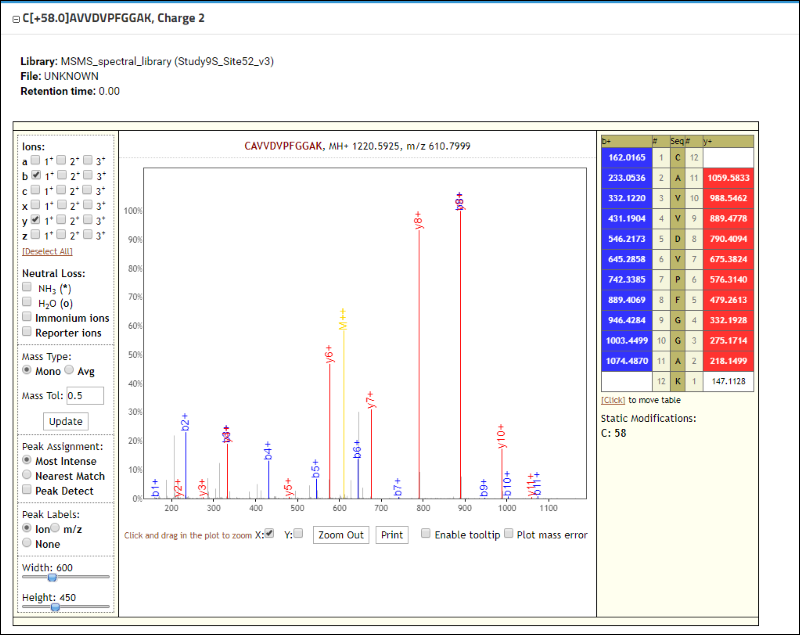
Scroll down further on the peptide details page to see an interactive spectrum viewer displaying the MS/MS spectrum for
CAVVDVPFGGAK from the spectrum library associated with this Skyline document. The name of the library is displayed in the title bar of the viewer. The viewer allows you to zoom in by clicking and dragging your mouse over the spectrum. You can also customize the view by selecting the ion types and charge states you want to display from the options menu to the left of the spectrum. A color-coded table to the right of the spectrum lists all the fragment ion m/z values for the peptide, with colored boxed representing the matching fragment ions found in the library spectrum. These spectral library views can be used to support publications in peer-reviewed journals or share data with collaborators.
Customizing Grid Views in Panorama
On your web-browser click the back button, or scroll to the top of the page and click on the ZIP file name in the navigation trail just under your folder name as shown in the image below. This will take you back to the documents details page.
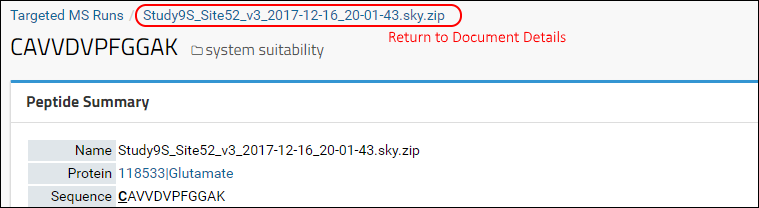
LabKey Server, on top of which Panorama is built, allows customizations of all tabular views. You can select which columns you would like to see, order them according to your preference, and define a sorting order. The
Precursor List table on this page has blank annotation columns since the researchers did not define any custom peptide or precursor annotations in this Skyline document.
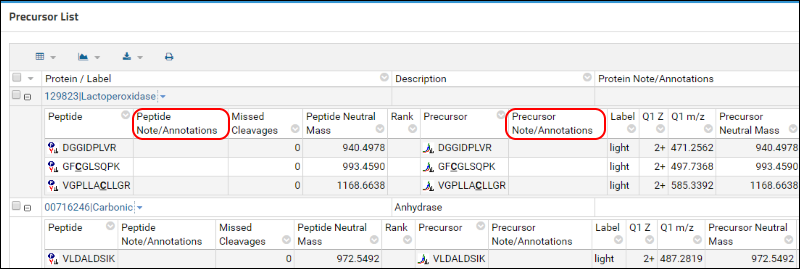
To hide these columns from the default view preform the following steps:
- Select (Grid Views) just under the Precursor List heading.
- Choose Customize Grid from the drop-down menu.
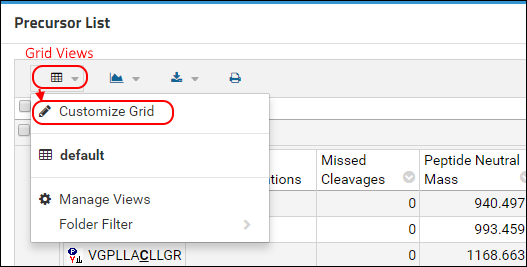
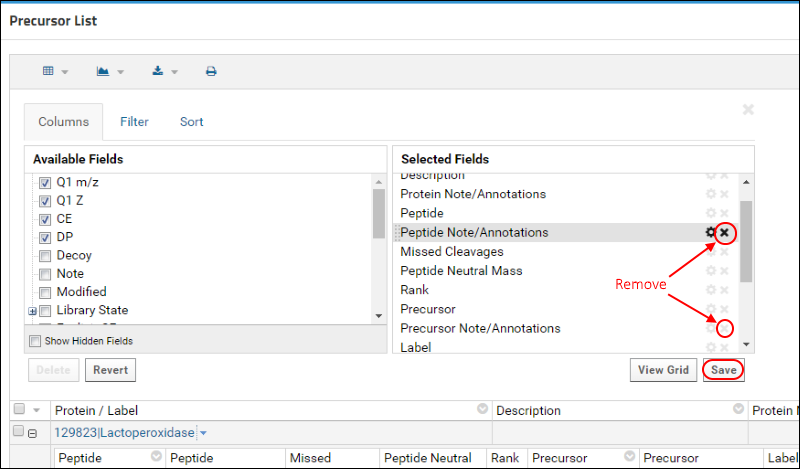
In the customization form that appears, you will see the Selected Fields panel on the right that shows the fields that are currently displayed.
- Hover over "Peptide Note/Annotations" below Peptide in the Selected Fields and click the (remove) icon.
- Do the same for "Precursor Note/Annotations" below Precursor.
- Click Save.
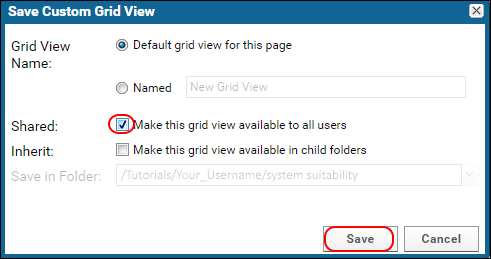
- In the popup, leave "Default grid view for this page" checked.
- Check "Make this grid view available to all users". This will cause all users with access to this folder to see the same table columns.
- Click Save.
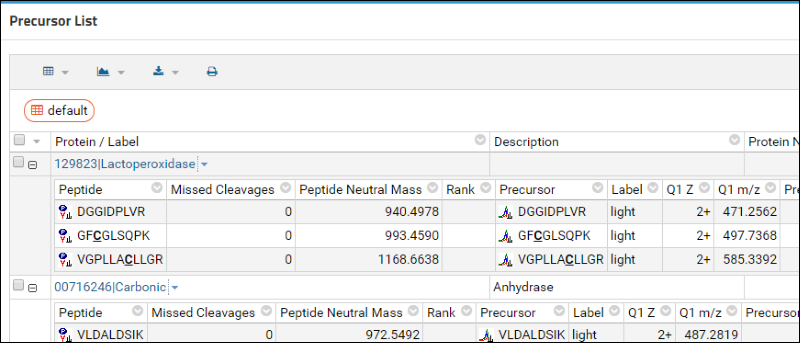
The annotation columns should no longer be displayed, as shown below:
You can view the LabKey Server documentation on customizing grid views (
https://www.labkey.org/wiki/home/Documentation/page.view?name=customViews) for more information.
Downloading Skyline Files from Panorama
Scroll up, if you need to, to view the
Document Summary box. You can download the original Skyline ZIP archive that was published to Panorama by clicking on the
Download link next to the file name.
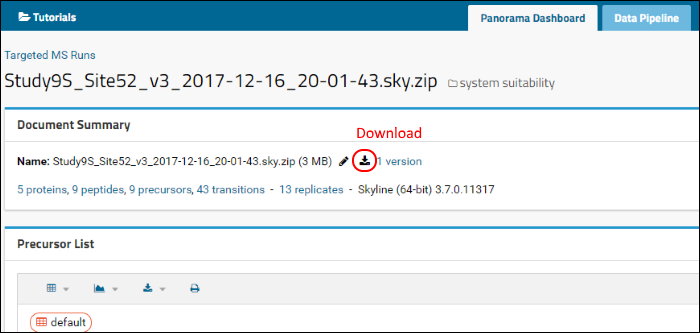
To open this zip file in Skyline do the following:
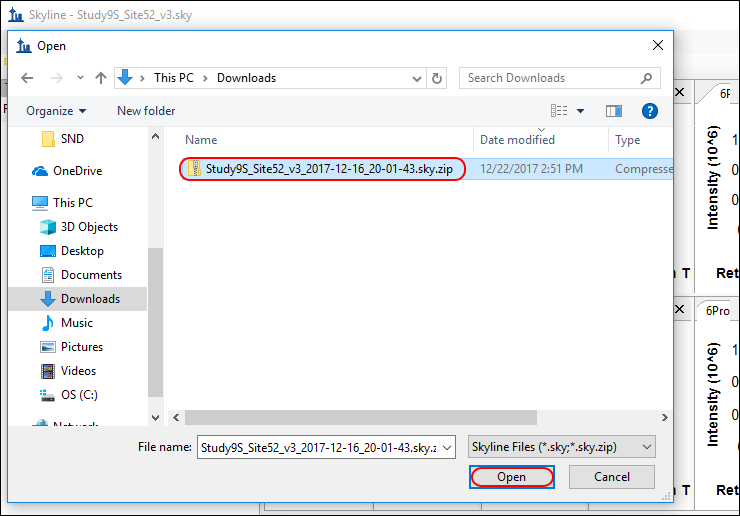
- On the File menu, click Open (Ctrl-O).
- Navigate to and select the zip file that you downloaded.
- Click the Open button.
Publishing to Panorama provides a convenient way to share Skyline documents with other researchers. You will see later in this tutorial how to give other users access to the folders you create in Panorama.
Uploading Documents Using the Panorama Web Interface
Using the Skyline
 Publish to Panorama
Publish to Panorama toolbar button or menu item is the most convenient way to get your documents into Panorama. But this can also be done by using the Panorama web interface. Next, you will use the web-interface to upload another document to the "system suitability" folder.
- In Skyline, open the file "Study9S_Site54_v2.sky".
This file contains results acquired at another site in the CPTAC system suitability study. Prepare this file for upload to Panorama by doing the following:
- In the File menu, click Share.
- Click the Minimal button in the form that appears.
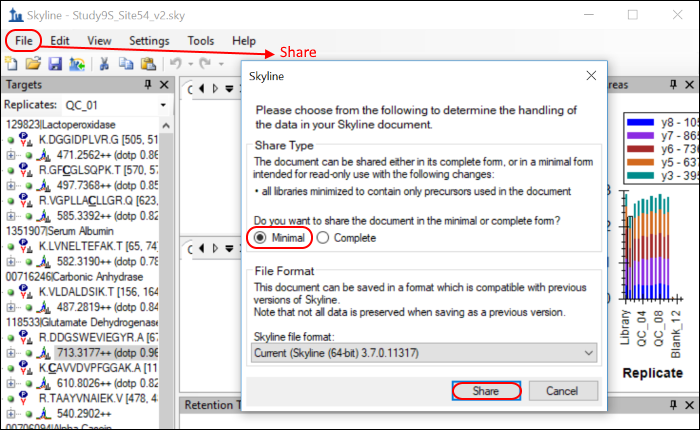
- Save the ZIP file on your computer using the Share Document form presented by Skyline.
- In Panorama, go to the "system suitability" folder.
- Click the Data Pipeline tab in the upper right corner.
- Click Process and Import Data as shown below.

- In the Files browser, click Upload Files.
- Click Browse or Choose File.
- Select "Study9S_Site54_v2.zip" that you just saved through Skyline.
- In the Description field enter "Site 54".
- Click Upload.
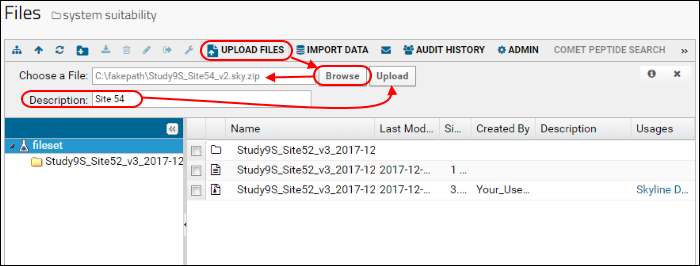
Once the file has been uploaded it will appear in the list of files in your folder.
- Check the box next to Study9S_Site54_v2.zip.
- Click Import Data in the toolbar.

- Confirm that Import Skyline Results is selected.
- Click Import in the Import Data popup.
Wait for the file import to complete and then click the
Panorama Dashboard tab. You will now see 2 files (from sites 52 and 54) available under
Targeted MS Runs.
Search Features in Panorama
Panorama allows you to search for proteins, peptides and even peptides with specific modifications across all the documents contained in a folder and its sub-folders. These search features are particularly useful for large projects and data sets, and help to compare results between data acquired over a long period of time.
On the "system suitability" folder page look at the
Mass Spec Search box. You will see three available search options under the tabs
Protein Search, Peptide Search and Modification Search. You will use the modification search option later in the tutorial. To perform a peptide search do the following:
- Click the Peptide Search tab.
- Enter "GFCGLSQPK" in the Peptide sequence field.
- Check Exact matches only.
- Click the Search button.
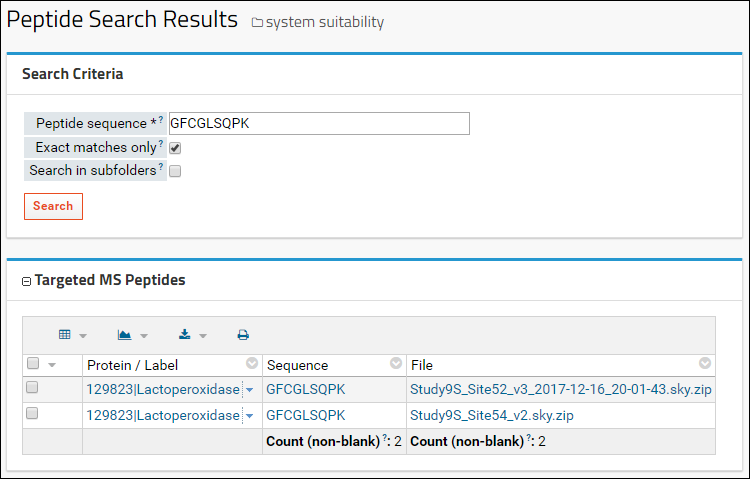
On the results page, in the
Targeted MS Peptides box, you will see a list of documents in your folder where this peptide was observed. This peptide was found in both documents uploaded (sites 52 and 54) to this folder. You can view the details for the peptide in a document by clicking on the peptide sequence.
Sharing Data in Panorama
Panorama allows you to set fine-grained access control on your folders. You can grant access to your folders to individual collaborators or groups of researchers, and you can also configure the type of access (read-only, read and edit etc.).
To change the permissions on the "system suitability" folder, first, make sure that you are on the "system suitability" folder page. If not, return by hovering over your project name (e.g. Tutorials) in the menu bar below the Panorama icon, opening the project and folders, then clicking your "system suitability" folder as shown in the image below.
To change the permissions on this folder, navigate to the
Permissions page by doing the following:
- On the menu bar, choose (Admin) > Folder > Permissions.
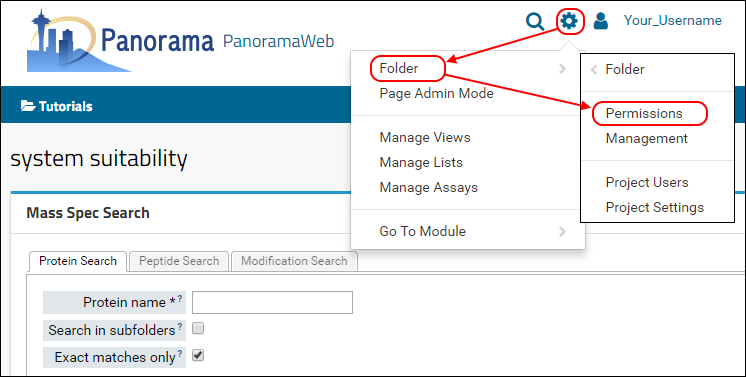
If you did not configure the permissions for this folder during the folder creation process, the permissions page should look like this.
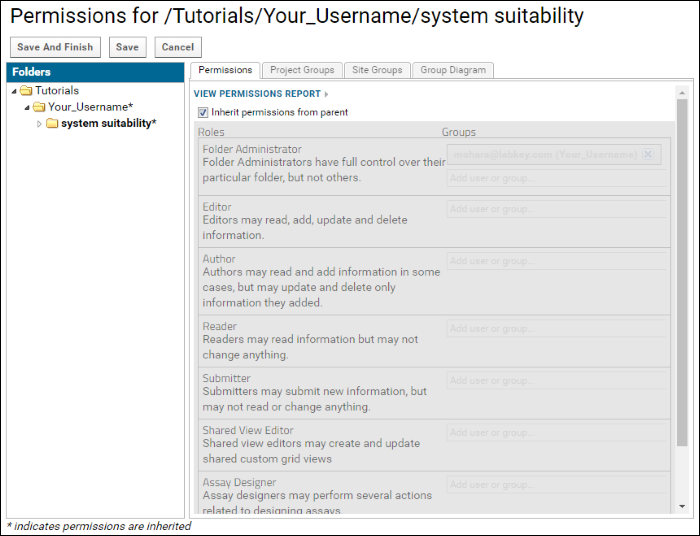
The email address selected for the
Folder Administrator role should be the email address with which you requested a project on panoramaweb.org. The Folder Administrator has full control over a folder. To add Brendan MacLean (Skyline team) to the Editor role, which will give him the ability to add and delete documents to this folder, follow these steps:
- Uncheck "Inherit permissions from parent", if it is checked.
- Select ‘brendanx@uw.edu (Brendan MacLean)’ from the combo-box next to Editor.
- Click Save.
To add Vagisha Sharma (Panorama team) to the Reader role:
- Select ‘vsharma@uw.edu’ from the combo-box next to Reader.
- Click Save.
The Reader role allows users to view data in a folder but they may not add or delete documents. The permissions page should now look like this:
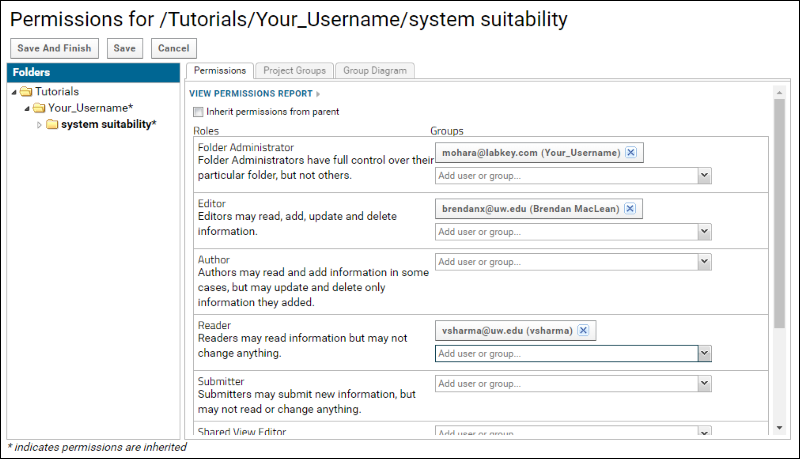
Instead of adding individual users to a role you can create a group of users and assign a role to the group. Only Project Administrators have permissions to create user groups. If you are working in your folder in the “Tutorials” project on panoramaweb.org, you do not have project administrator privileges. However, if you are working through this tutorial in your own project on panoramaweb.org or any other Panorama server, follow these steps to create a "Seattle collaborators" group and assign it to the
Reader role:
- On the Permissions page click on the Project Groups tab.
- In the New group name field enter "Seattle collaborators" and click Create New Group.
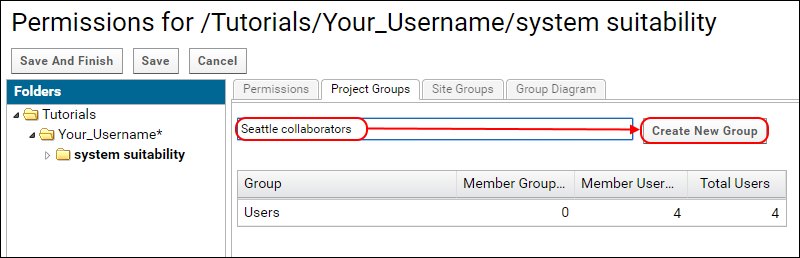
- In the popup select Brendan and Vagisha from the pull down list to add them to the group.
- Click Done.

- Select the Permissions tab.
- Add the new "Seattle collaborator" group to the Reader role.
- Click Save.
- Select your "system suitability" folder in the folder navigation column.
- Add the "Seattle collaborator" group to the Reader role for this folder.
- Click Save and Finish.
Members of the "Seattle collaborators" group will be able to view the "system suitability" folder if they have a direct link to the folder. They will not be able to view the parent folder (e.g. Your_Username) on the home page or navigate to the "system suitability" folder through the folder navigation UI. To access a folder from the UI, users or user groups require read permissions not only on the folder but also on the parent folders all the way up to the root project folder. To enable the "Seattle collaborators" group to access the "system suitability" folder via the navigation UI, assign the
Reader role to this group in the parent folder (e.g. Your_Username).
You can add more users later to the "Seattle collaborators" group. All users in this group will have read access to the "system suitability" folder. If you would like to add a user to the
"Seattle collaborators" group that does not already have an account on Panorama, you can do that by following these steps:
- Select (Admin) > Folder > Permissions.
- Click the Project Groups tab.
- Click the "Seattle collaborators" group.
- In the popup, click Manage Group.
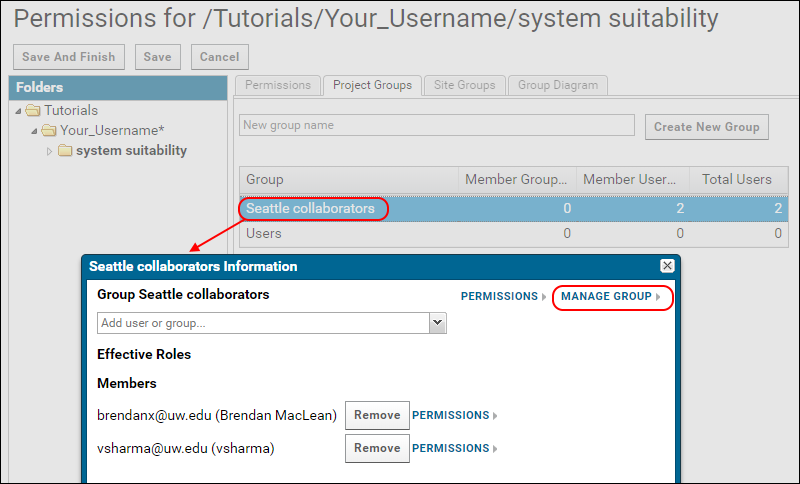
- On the Manage Group page, enter the email address of the new user in the Add New Members field.
- Scroll down and click Update Group Membership.
An email will be sent to the new user and they will be able login to view the data in the "System suitability" folder once they setup a password for their account.
You can view the LabKey Server documentation on configuring permissions (
https://www.labkey.org/wiki/home/Documentation/page.view?name=configuringPerms) for more information.
Viewing of PTM-containing Peptides in Panorama
Many journals require visual display of MS/MS spectra of peptides containing post-translational modifications such as phosphorylation or acetylation. You can make use of Skyline to build spectral libraries from your peptide search results and view the annotated spectra in the Skyline Spectral Library Explorer. However, using the resulting Skyline document to satisfy journal requirements requires that you send the Skyline document to reviewers, and that they have a local installation of Skyline to view the document. Alternatively, you can publish the documents to Panorama, where reviewers can view the spectra in the Panorama spectrum viewer.
Next, you will import a Skyline document to Panorama that contains phosphorylated peptides, search for modified peptides and view their spectra.
Create a new sub-folder in "Your_Username" folder called "Phospho peptides", using another method for creating sub-folders in Panorama than you used before:
- In the header bar, select (Admin) > Folder > Management.
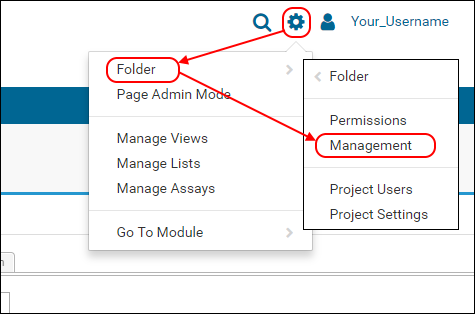
- On the Folder Management page, make sure that the Folder Tree tab is selected, and "Your_Username" folder is highlighted.
- Click Create Subfolder.

- In the Create Folder wizard enter Name: “Phospho peptides”.
- Select Folder Type, "Panorama" as before.
- Click Next.
- For Users/Permissions keep the default selection: Inherit from Parent Folder.
- Click Next.
- Select Experimental Data on the Configure Panorama Folder page.
- Click Finish.
To add to the new folder a Skyline document containing a spectral library from a phosphorylation experiment, perform the following steps:
- In Skyline, open the file "Phospho_0921_2010_Thr99_v1_P only.sky" from the tutorial folder.
- Click the
 Publish to Panorama toolbar button.
Publish to Panorama toolbar button.
- Select the “Phospho peptides” folder from the folder tree in the Publish Document dialog.
- Click OK.
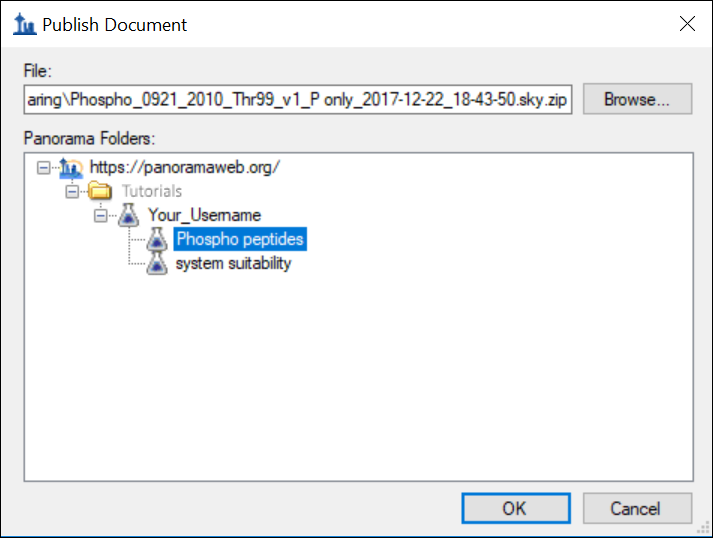
Wait for data import to complete and then go back to Panorama in your web-browser, and navigate to the “Phospho peptides” folder. Refresh the page (F5 in most browsers) if you are already on the “Phospho peptides” folder page. You should see the file that you just imported to Panorama in the
Targeted MS Runs table. Click on the name of the file to see the document details. Clicking on any of the peptide sequences will take you to the peptide details page where you can view the library spectrum for the peptide. Click the back button to go back to the folder page, or use the
Phospho peptides link.
To find all peptides in this document containing a phosphorylation, perform the following steps:
- In the Mass Spec Search box, select the Modification Search tab.
- Choose the Delta Mass option for Search By, if it is not already selected.
- Enter "S" in the Amino Acid field.
- Enter "80" in the Delta Mass field.
- Click Search.
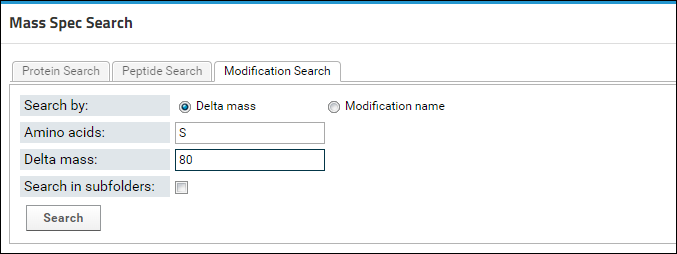
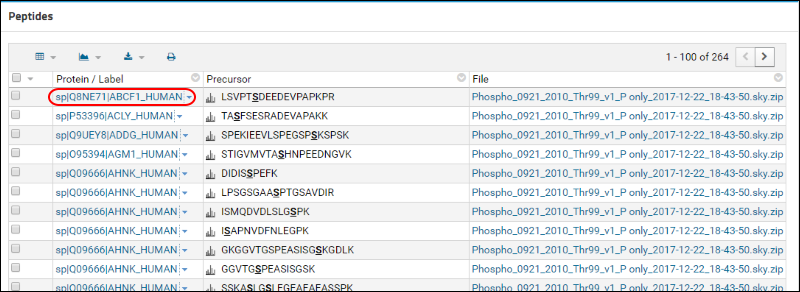
You will now see a list of all the serine phosphorylated peptides in the document. Click on the first peptide in the list, LSVPTSDEEDEVPAPKPR, to view its library spectrum. Click back in your browser. In the
Peptide Summary box click on the name of the protein (sp|Q8NE71|ABCF1_HUMAN) to which this peptide belongs.
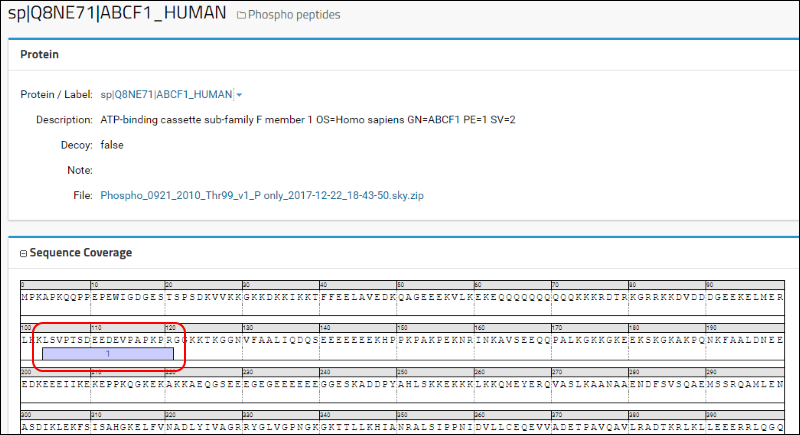
You will be taken to the protein details page where you can see the location of this peptide in the protein sequence.
Conclusion
In this tutorial, you have been introduced to new ways you can share Skyline documents with collaborators, reviewers and the general public using Panorama, and the PanoramaWeb.org web site hosted by the MacCoss Lab at the University of Washington. You have requested a project on PanoramaWeb, registered an administrator account for that project with your email address, and created 2 new folders in that project. You have explored data from Skyline documents in these folders, searched for specific peptides and post-translational modifications, and customized access to the data in them. If you worked on this tutorial in your own project instead of the "Tutorials" project, you can now just as easily go back to the folder management form and delete these tutorial folders to return your new project to its original state, ready for your own experimental data.
We wish you luck in your research and hope that you find these tools useful in sharing and disseminating your findings.