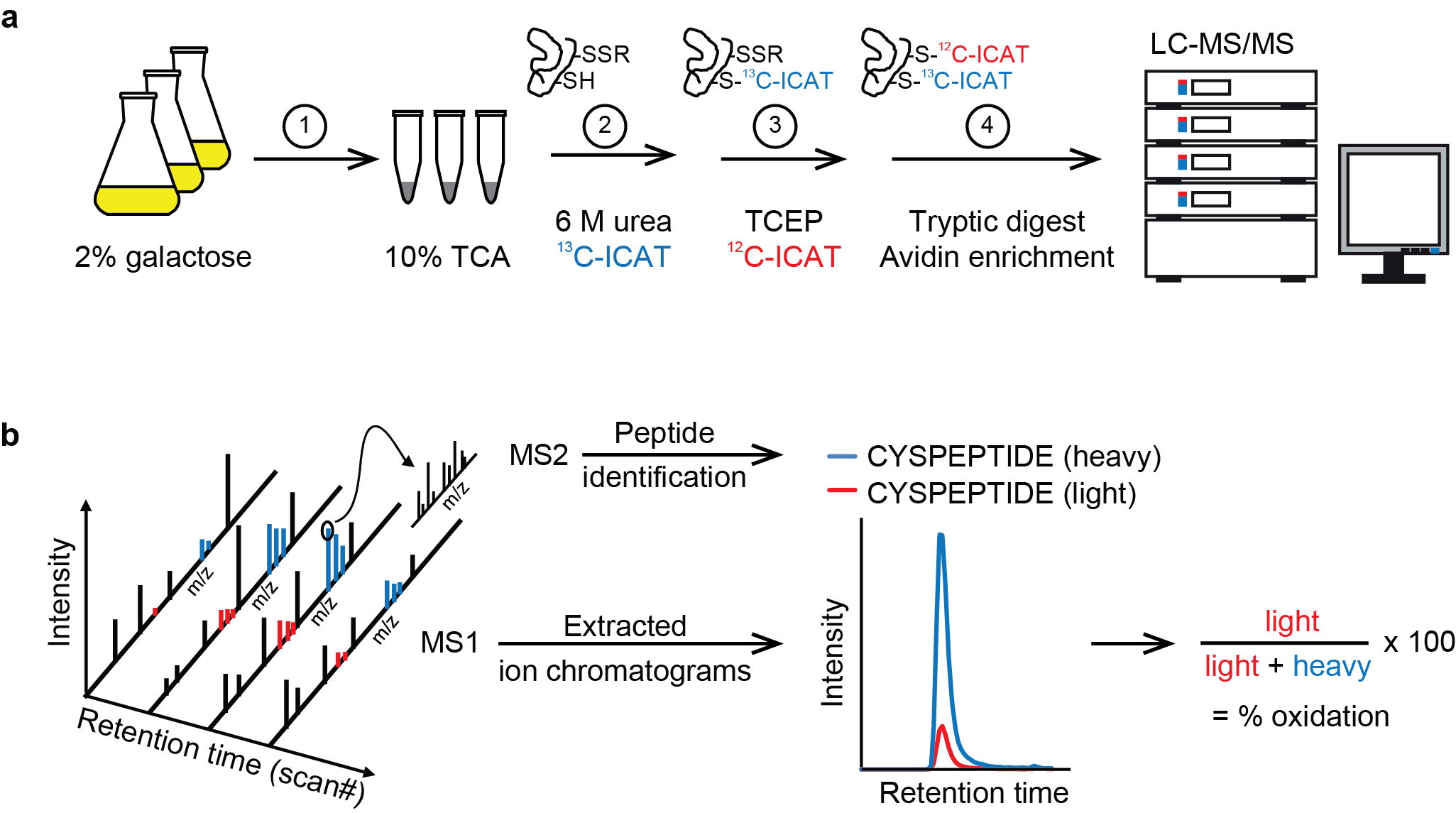
Site-resolved, large-scale analysis of the yeast redoxome. Yeast cells grown in 2% galactose medium were immediately frozen in 10% TCA (1). Extracted proteins were denatured in 6 M urea in the presence of heavy ICAT (13C-ICAT) to label free thiol groups (2). Reversibly oxidized cysteine residues were reduced by TCEP and labeled by light ICAT (12C-ICAT) (3). Proteins were digested with trypsin and ICAT-labeled peptides were enriched by streptavidin affinity chromatography (4). Following cleavage of the biotin tag, peptides were analyzed in duplicate by LC-MS/MS. (b) For the accurate determination of the redox state of cysteine residues, peptides were identified based on fragment ions observed in MS2 spectra using MaxQuant (v.1.4.1.2) and then quantified by extracting MS1 ion chromatograms using Skyline (v.2.5.0). Following integration of peak areas of heavy and light peptide variants, the proportion of reversibly oxidized cysteine residues (% oxidation) was calculated.
Please note: When the retention time profiles of “light” and “heavy” peaks were inconsistent, the respective XICs were excluded from quantification. The information which XICs were not quantified can be found in the tables below (denoted by “nq”).
Table S2. Oxidation status of cysteine containing peptides of non-stressed yeast cells. MS1 ion chromatograms were extracted using Skyline (v.2.5.0). Integrated peak areas of the “heavy” (13C-ICAT labelled) and “light” (12C-ICAT labelled) version of the peptide ions were exported and the proportion of reversibly oxidized (12C-ICAT labelled) cysteine residues (%oxidation) was calculated. The table contains all cysteine containing peptide sequences quantified in Skyline in at least two of three biological replicates.
| Attached Files | ||
Table S3. Peptides identified and quantified in oxICAT labeled isolated mitochondria. The table contains all cys containing peptide sequences identified with posterior error probability (PEP) < 0.01 and intensity >0 by MaxQuant search. Reverse entries were removed.
| Attached Files | ||
Table S5. Oxidation status of cysteine containing peptides of yeast cells upon H2O2 stress. This table contains all cysteine containing peptide sequences quantified in Skyline in ≥ 2 biological replicates of H2O2 treated samples and ≥ 2 biological replicates of control samples.
| Attached Files | ||
Peptides identified in OxICAT experiments of mia40-4int and wildtype control samples. This table contains all cysteine-containing peptide sequences identified in mia40-4int samples wildtype control samples with posterior error probability (PEP) < 0.01 and intensity > 0 and all non-cysteine containing peptide sequences identified with PEP<0.01. It includes quantification data for all peptide sequences quantified in Skyline in ≥ 3 biological replicates of mia40-4int samples and in ≥ 3 biological replicates of wildtype control.
| Attached Files | ||