Pino LK, Searle BC, Huang EL, Noble WS, Hoofnagle AN, MacCoss MJ. Calibration Using a Single-Point External Reference Material Harmonizes Quantitative Mass Spectrometry Proteomics Data between Platforms and Laboratories. Analytical Chemistry. 2018 Oct 11;90(21):13112-7.
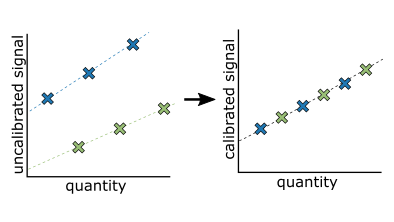
Mass spectrometry (MS) measurements are not inherently calibrated. Researchers use various calibration methods to assign meaning to arbitrary signal intensities and improve precision. Internal calibration (IC) methods use internal standards (IS) such as synthesized or recombinant proteins or peptides to calibrate MS measurements by comparing endogenous analyte signal to the signal from known IS concentrations spiked into the same sample. However, recent work suggests that using IS as IC introduces quantitative biases that affect comparison across studies due to the inability of IS to capture all sources of variation present throughout an MS workflow. Here we describe a single-point external calibration (EC) strategy to calibrate signal intensity measurements to a common reference material, placing MS measurements on the same scale and harmonizing signal intensities between instruments, acquisition methods, and sites. We demonstrate data harmonization between laboratories and methodologies using this generalizable approach.
Sample preparation. The data regenerated in this work used yeast strain BY4741 (MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0) (Dharmacon) cultured in YEPD to mid log phase, then treated with NaCl to a final concentration of 0.4M NaCl. Cell pellets were harvested and lysed individually with 8M urea buffer solution and bead beating (7 cycles of 4 minutes beating with 1 min rest on ice). Cell lysates were reduced, alkylated, digested for 16 hours, and desalted with a mixed-mode (MCX) method.
Selected reaction monitoring mass spectrometry (SRM-MS). Data were acquired using selected reaction monitoring (SRM) on a Proxeon EasyLC coupled to a Thermo Altis triple quadrupole mass spectrometer. Peptides were separated by reverse phase liquid chromatography using pulled tip columns created from 75 um µm inner diameter fused silica capillary (New Objectives, Woburn, MA) in-house using a laser pulling device and packed with 3 μm ReproSil-Pur C18 beads (Dr. Maisch GmbH, Ammerbuch, Germany) to 30 cm. Trap columns were created from 150 um µm inner diameter fused silica capillary fritted with Kasil on one end and packed with the same C18 beads to 3 cm. Solvent A was 0.1% formic acid in water (v/v), solvent B was 0.1% formic acid in 80% acetonitrile (v/v). For each injection, approximately 1 μg total protein was loaded and eluted using a 90-minute gradient from 5 to 40% B in 25 minutes, 40 to 60% B in 5 minutes, followed by a 15 minute15-minute wash and then 15 minutes equilibration back to initial conditions. Total analytical run time was 45 minutes. Thermo RAW files were imported into Skyline1110 (Skyline-daily version 4.1.1.18151) for processing and Total Area Fragment results were exported using a Custom Report.
Data independent acquisition mass spectrometry (DIA-MS). Data were acquired using data-independent acquisition (DIA) on a Waters NanoAcquity UPLC coupled to a Thermo Q-Exactive HF orbitrap mass spectrometer. Peptides were separated by reverse phase liquid chromatography using pulled tip columns created from 75 um µm inner diameter fused silica capillary (New Objectives, Woburn, MA) in-house using a laser pulling device and packed with 3 μm ReproSil-Pur C18 beads (Dr. Maisch GmbH, Ammerbuch, Germany) to 30 cm. Trap columns were created from 150 um inner diameter fused silica capillary fritted with Kasil on one end and packed with the same C18 beads to 3 cm. Solvent A was 0.1% formic acid in water (v/v), solvent B was 0.1% formic acid in 98% acetonitrile (v/v). For each injection, approximately 1 μg total protein was loaded and eluted using a 90-minute separating gradient starting at from 5 and increasing to 35% B, followed by a 40 minute40-minute wash and equilibration (total 130 minute method). DIA methods followed the chromatogram library workflow, described in greater detail elsewhere1211. Briefly, the untreated (reference) samples and osmotic shocked peptide samples were pooled 1:0.33:0.33:0.33 to create a library sample, and a Thermo Q-Exactive HF was configured to acquire six gas phase fractions, each with 4 m/z DIA spectra using an overlapping window pattern from narrow mass ranges. For quantitative samples, the Thermo Q-Exactive HF was configured to acquire 25x 24 m/z DIA spectra using an overlapping window pattern from 388.43 to 1012.70 m/z. The specific windowing schemes for both the chromatogram library construction and quantitative experiments are described in Supplemental Table 1. All DIA spectra were programmed with a normalized collision energy of 27 and an assumed charge state of +2.
Thermo RAW files were converted to .mzML format using the ProteoWizard package (version 3.0.10106), where they were peak pickedcentroided using vendor provided file reading libraries. Converted acquisition files were processed using EncyclopeDIA (version 0.7.0) configured with default settings (10 ppm precursor and, fragment, and library tolerances, considering both B andonly Y ions, and trypsin digestion was assumed). EncyclopeDIA was configured to use EncyclopeDIA features were submitted to Percolator (version 3.1) for validation at 1% FDR Percolator (version 3.1).