Table of Contents |
guest 2024-07-04 |
Neat Unscheduled Runs
Plasma Wide Window Scheduled
Plasma Narrow Windows Scheduled
Absolute Quantitation - PQ500
Biognosys PQ500 Introduction
This tutorial will show you how to create a targeted MS2 assay that uses heavy standards for absolute quantitation. The Biognosys PQ500 standard is used as the source of heavy standards. We used the Vanquish Neo LC, ES906A column and a trap-and-elute injection scheme with a 60 SPD method and a 100 SPD method. The gradients have been designed so that compounds elute over a large portion of the experiment spans.

Setting up the Skyline Document
Pierce retention time calibration mixture (PRTC) is used here to create an indexed retention time (iRT) calculator. Along with a spectral library, the iRT calculator will aid Skyline in picking the correct LC peaks in the steps that follow. See the Skyline iRT tutorial for more details. Here we will use an iRT calculator created with Koina. After setting up the LC and column, we run unscheduled PRTC injections to ensure that the LC and MS system is stable. The method file 60SPD_PRTC_Unscheduled.meth can be used for this. The prtc_unscheduled.csv file could be used to import into a tMSn table if making a method from scratch. We like to use Auto QC with Panorama to store all our files, and to automatically upload and visualize QC data.
Now we will create a Skyline document for analyzing PQ500 heavy labeled peptides. Biognosys supplies a transition list with intensities and iRT values that can be used to create a spectral library and iRT calculator. We'll show you though also how you can use use Koina integration with Skyline to create a spectral library and iRT library from a list of peptides sequences if you don't know anything about them. We also tend to like to use Koina spectral libraries even if supplied lists of transitions, because we will be using PRM Conductor to automatically filter the transitions. At the end of this section, we’ll be ready to perform unscheduled PRM for the PQ500 heavy-labeled peptide standards.
Transition Settings
Open up Skyline Daily and create a new document. Save the document as Step 1. Setup Skyline Documents/pq500_60spd_neat_multireplicate.sky.
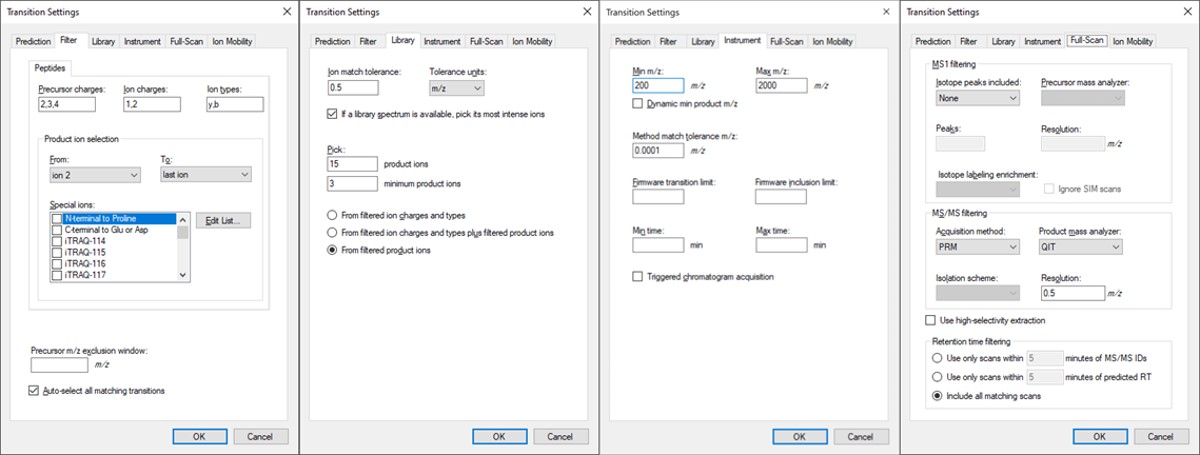
- Open Settings / Transition Settings / Filter. Set Precursor charges to ‘2,3,4’, Ion Charges to ‘1,2’, and Ion types to ‘y,b’. In the Product ion selection section, select ‘From ion 2’ to ‘last ion’. Un-select N-terminal to Proline and keep “Auto-select all matching transitions” checked.
- In the Library tab, set Ion match tolerance 0.5, check the box “If a library spectrum…”, set Pick 15 product ions with minimum 3 product ions. 15 is a large number, but we will refine the transitions later with the PRM Conductor tool. Select “From filtered product ions”.
- In the Instrument tab, set Min m/z 200, and Max m/z 2000. Set the Method match tolerance m/z to 0.0001. This helps Skyline to differentiate between precursors that have very close m/z. As we’ll see later, there are sometimes still peptides with different sequences but the same exact m/z.
- In the Full-Scan tab, set MS1 filtering / Isotope peaks included to None. If there are any precursor transitions, PRM Conductor will think that the document is in DDA mode, and will not be full featured. Set MS/MS filtering / Acquisition Method to PRM, Product mass analyzer to QIT, and Resolution to 0.5 m/z. Set Retention time filtering to Include all matching scans. Press Okay to close the Transition Settings.
Peptide Settings
Open Settings / Peptide Settings.
- In the Library tab, uncheck or remove any libraries that are there, for simplicity's sake.
- In the Modifications tab, make sure that Carbamidomethyl (Cysteine) and Oxidation (Methionine) modifications are enabled. The C-term R and C-term K isotope modifications should be enabled. Isotope label type and Internal standard type should be set to heavy.

iRT Calculator from PRTC
- In the Prediction tab, select the calculator icon and press Add. Add a name like PRTC. In the iRT Standards drop-down menu, select Pierce (iRT-C18). Press the Create button, and select Yes when asked if you want to create a new database file. Give the file a name like prtc.irtdb. Press Okay.
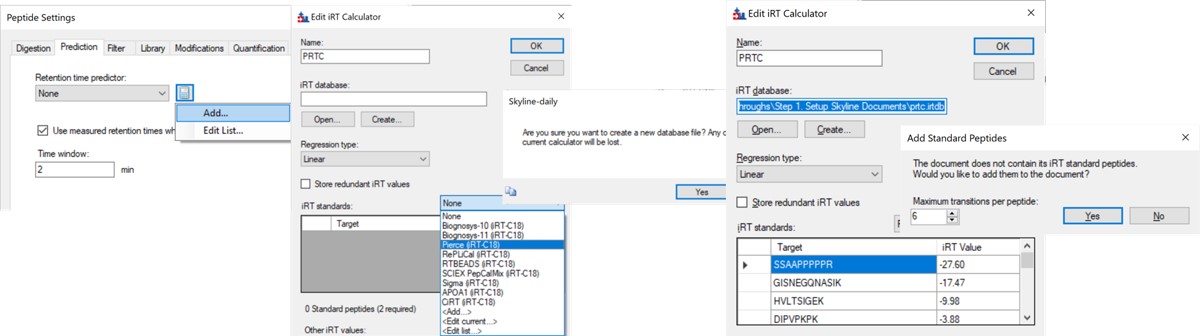
- Go back into the Peptide Settings / Prediction tab, click the dropdown arrow and select Edit current. Set the window to 2 minutes, and press Okay.
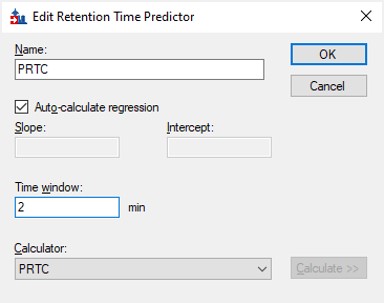
Importing the Transition List
Use File / Import / Transition List and select the file Step 1. Setup Skyline Documents/biognosis_pq500_transition_list.csv. A dialog opens that shows the mapping of the file headers to Skyline variable names. Press Okay to continue on. A new dialog prompts us that 624 transitions are not recognized. These are water losses that we don't necessarily need. We could define water loss transitions in the Settings tabs if we really wanted them. Press Okay twice to exit the iRT calculator dialogs. A new dialog will ask if you want to add the Standard Peptides, choose 6 transitions and press Yes.
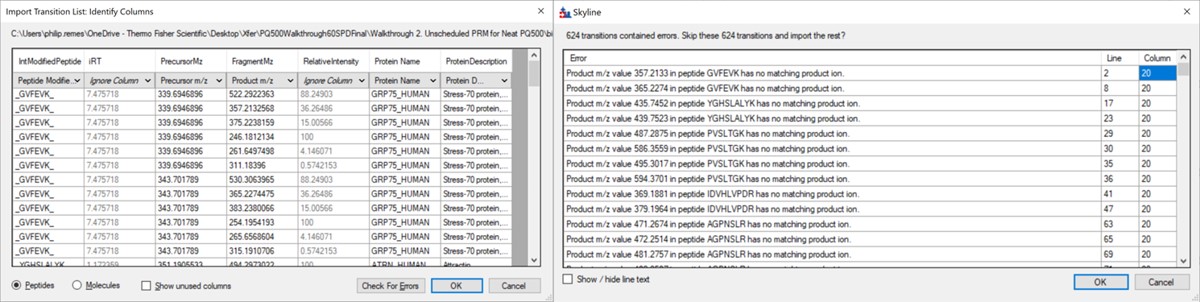
Another dialog appears, asking if we want to make an iRT calculator. The Biognosys values are presumably based on experiment, and are slightly more accurate than the in silico predicted iRT values from Koina, so Click Create. You'll be asked if you want to create a spectral library from the intensities in the transition list. Feel free to press Create if you want, but we will press Skip and use Koina to predict the intensities next. The Skyline document will update and in the bottom right border will be displayed 579 prot, 818 pep, 1622 prec, 9020 tran.

Koina
Now we’ll generate a spectral library with Koina.
- Open Tools / Options and go to the Koina tab. Set the intensity model to ms2pip_2021_HCD, the iRT model to AlphaPept_rt_generic, and NCE to 30. Note that our original assay was created when this option tab said Prosit, but it had changed in a new Skyline Daily release by the time that this walkthrough was created. Press Okay.
- Go to Settings / Peptide Settings / Library / Create and select Data source Koina, NCE 30, and set a name and output path for the library. Press Finish. Make sure the library is selected in Peptide Settings Library and press Okay. If everything works correctly and you have internet access, your computer will communicate with the Koina server and return the spectral library to you, and the peptides in the Skyline tree will receive little spectra images to the left of the sequences, like below.
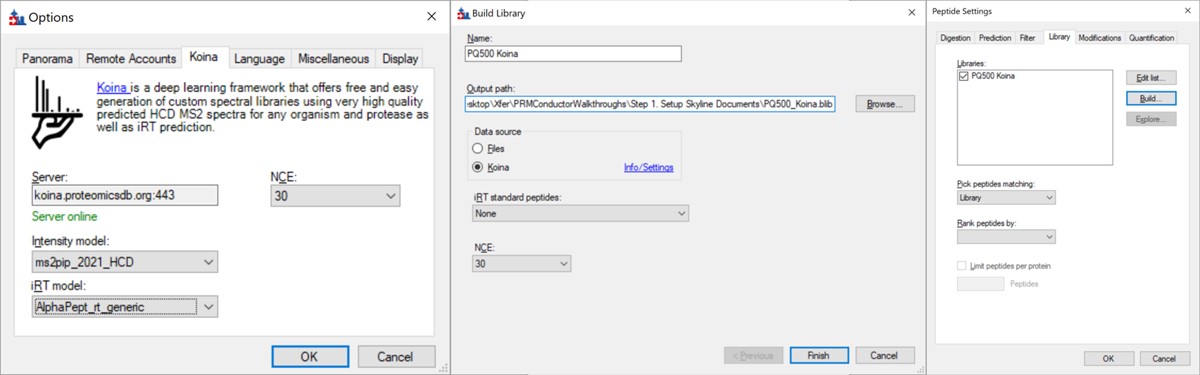
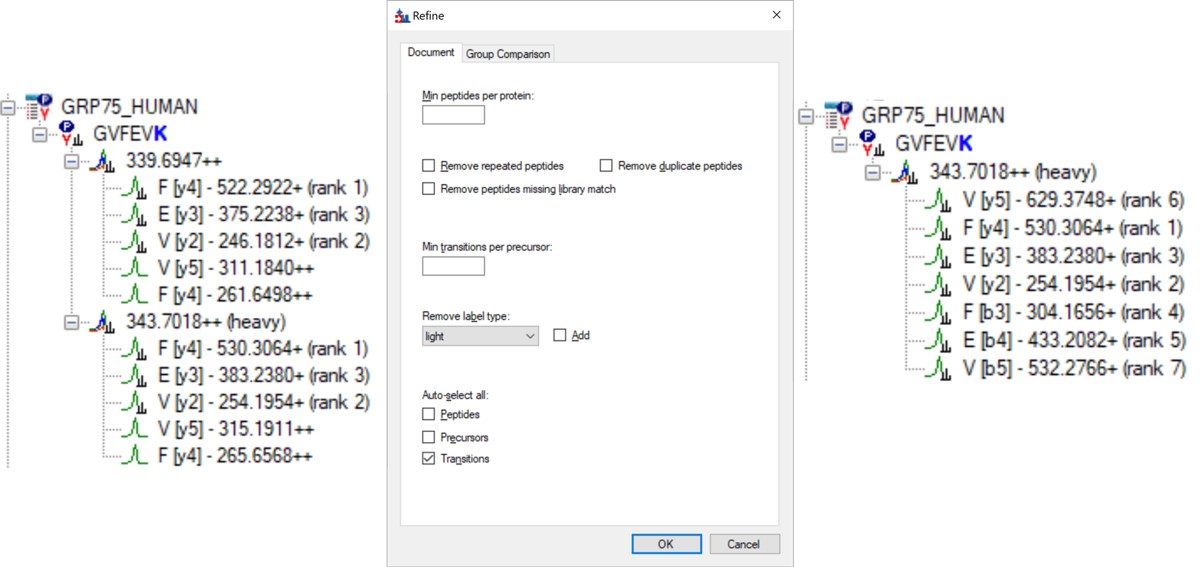
- Open Refine / Advanced and select Remove label type light, and click the box for Auto-select all Transitions, and press Okay. If you expand a peptide, like the first one, GVFEVK, it will have up to 15 transitions, instead of the up to 6 in the Biognosys transition file. Save the Skyline document. This is the end of Step 1: Setting up the Skyline Document.
Neat Unscheduled Runs
- Unscheduled PRM for Neat Heavy PQ500 with Multiple Replicates
Unscheduled PRM for Neat Heavy PQ500 with Multiple Replicates
In this step, we’ll use Skyline to create a set of unscheduled PRM methods for the 804 PQ500 peptides. At the end of this step, we will have created 10 Unscheduled PRM methods for both 60 and 100 SPD, acquired data for them, loaded the results into Skyline, and assessed the results. We’ll be ready to look at our standards spiked into matrix in the next step.
Alternative DIA GPF Acquisition
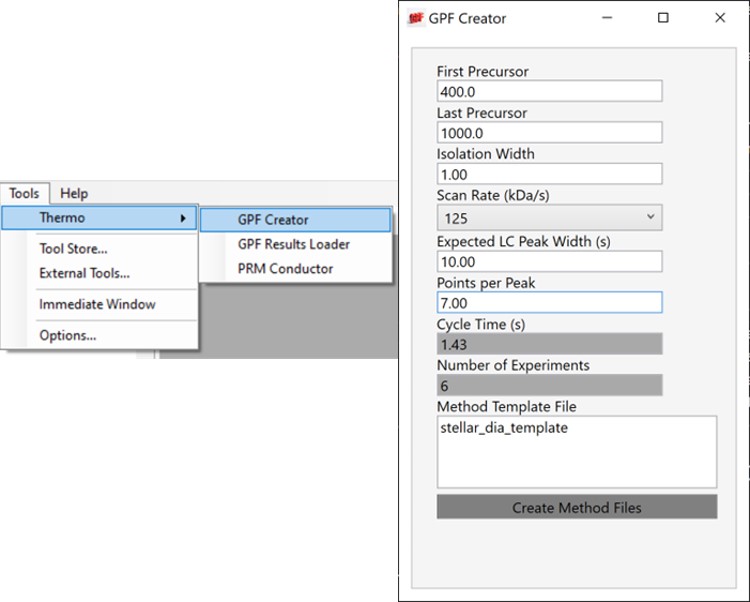
Alternatively, especially as the number of heavy peptides increases, one could opt to use data independent acquisition (DIA) of the neat, heavy standards to find their retention times. One would simply find the smallest and largest m/z of the peptides in question and use the Thermo method editor to create a DIA method. For example, we have had success in some neat standard cases using a single injection with 4 Th isolation width. However multiple gas-phase fractions (GPF) could be acquired with narrower isolation widths, as in the technique we use for identification of unknowns. We included a little helper application in our Thermo suite of external tools called GPF creator that spawns GPF instrument methods. Given a set of parameters, namely a precursor m/z range and a Stellar DIA method template, it will create a cloned set of methods with the appropriate Precursor m/z range filled in. In the case below with Precursor m/z range 400-1000 and 6 experiments, methods would be created for the ranges 400-500, 500-600, all the way to 900-1000. The resulting .raw files could be used in the much the same way that we’ll use the unscheduled PRM data files in the coming steps, only that we would have to configure the Skyline Transition / Full Scan / Acquisition to DIA with the appropriate window scheme (Ex. 400 to 1000 by 1 with Window Optimization On). As it is, we continue on, using the Unscheduled PRM technique.
Creating Unscheduled PRM Methods
The newest Skyline release supports Stellar for exporting isolation lists and whole methods, which is convenient as it saves the step of importing isolation lists for each of the methods. However, for completeness we'll also describe how to use the isolation list dialog with manual import into method files.
Isolation List with Manual Import to Instrument Method Files
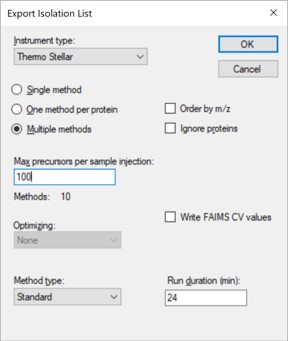
- Open the dialog using File / Export / Isolation List. Set Instrument type Thermo Stellar, select Multiple methods, Max precursors per sample injection to 100, Method type Standard. Note that the max value of 100 is approximate. A smaller number could be required for more narrow peaks or shorter gradients. Press Okay.
- A Save-As dialog will open. Use the name pq500_neat_unscheduled in the folder Step 2. Neat Unscheduled Multireplicates. In a moment, 10 .csv files with the suffix _0001 to _0010 will appear in the folder. An important aspect of the iRT workflow is that Skyline included the PRTC compounds in each of the 10 files. This allows Skyline to calculate the relative positions of the rest of the peaks, which will eventually be added to our iRT library.
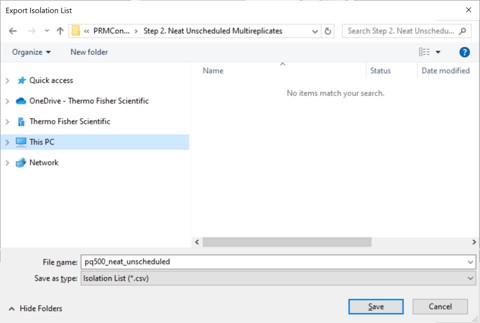
- Open the Stellar method editor and open the file Step 2. Neat Unscheduled Multireplicates/pq500_60spd_neat_multireplicate.meth. In the tMSn experiment in the bottom right you can find the Mass List table and press the Import button to load the first of the created isolation lists, pq500_neat_unscheduled_0001.csv. Use File / Save-As to save the method as pq500_neat_unscheduled_0001.meth. Do this for each of the other 9 isolation lists, creating a total of 10 methods. At this point you would create an Xcalibur sequence with these 10 methods, injecting an appropriate amount of neat standard (ex 10-100 fmol for a 1 ul/min, 24 min assay), and creating 10 raw files. We have performed this experiment for both 60 and 100 SPD methods, and put the resulting .raw files in the folder Step 2. Neat Unscheduled Multireplicates/Raw.

Skyline Method Export
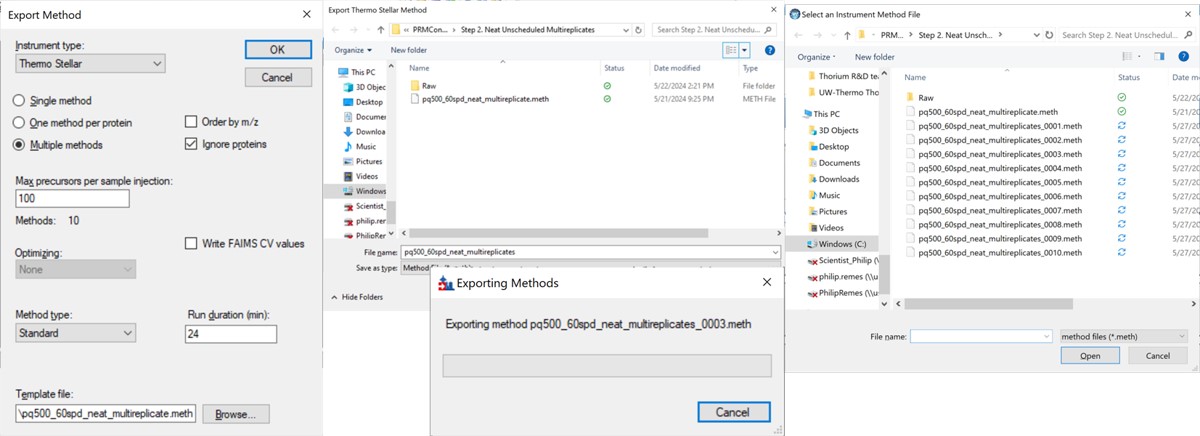
The more convenient way to create the unscheduled replicates is to use the Skyline File/Export/Method functionality. In the Export Method dialog, select Instrument type Thermo Stellar. Select Multiple methods, with Max precursors per sample injection 100. This is a ballpark number that has given enough points per peak for identification purposes for neat standards for a variety of experiment lengths. Click the Browse button and choose the pq500_60spd_neat_multireplicate.meth file in the Step 2. Neat Unscheduled Multireplicates folder. Use a name like pq500_60spd_neat_multireplicates and press Save, and Skyline will present a progress dialog. When it finishes, 10 new methods will be created with suffix _0001 through _0010, as shown below.
Loading Unscheduled PRM Data
- Take the .sky file that we have created, Step 1. Setup Skyline Documents/pq500_60spd_neat_multireplicate.sky and use File / Save As to save a copy with the name Step 2. Neat Unscheduled Multireplicates/pq500_60spd_neat_multireplicate_results.sky. If you want to work along with the 100 SPD method as well, you can save another version of the file with the corresponding name.
- Use File / Import / Results and select Add one new replicate. Give it the name NeatMultiReplicate. Press Okay, and then select all the raw files in the folder Step 2. Neat Unscheduled Multireplicates\Raw\NeatUnscheduled60SPD and press Open. Skyline will load the results. When they are finished loading, Save the Skyline document. Do the same for the 100 SPD .raw files in their .sky file.

- Now we will inspect the results to see if any peaks have been chosen incorrectly. Use File / Save As to save the Skyline document as Step 2. Neat Unscheduled Multireplicates/pq500_60spd_neat_multireplicate_results_refined.sky (and one for the 100 SPD file).
- Use View / Retention Times / Regression / Score to Run. Right click the plot and select Plot / Residuals. Right click the plot and make sure that Calculator is set to the calculator that we created, PRTC.
- Right click on the plot and select Set Threshold and enter 0.99. If the plot does not update to have some of the blue dots turn pink, then right click the plot and select Plot / Correlation, and then switch back to Plot / Residuals.
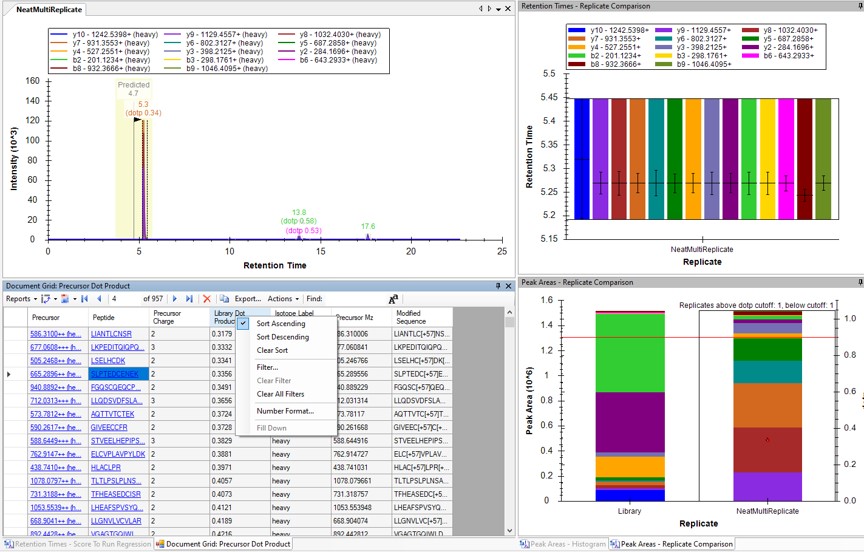
- Use View / Peak Areas / Replicates Comparison and setup the Skyline document something like in the figure below.
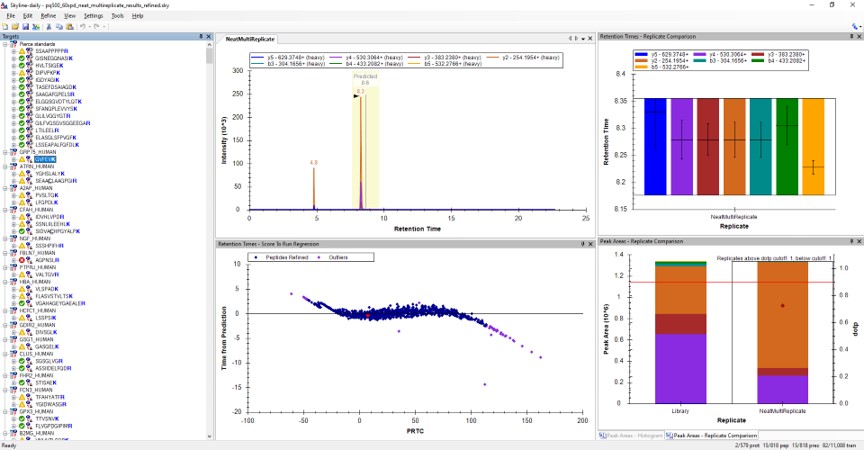
Reviewing the Unscheduled PRM Results
Using Score-to-Run RT Outliers
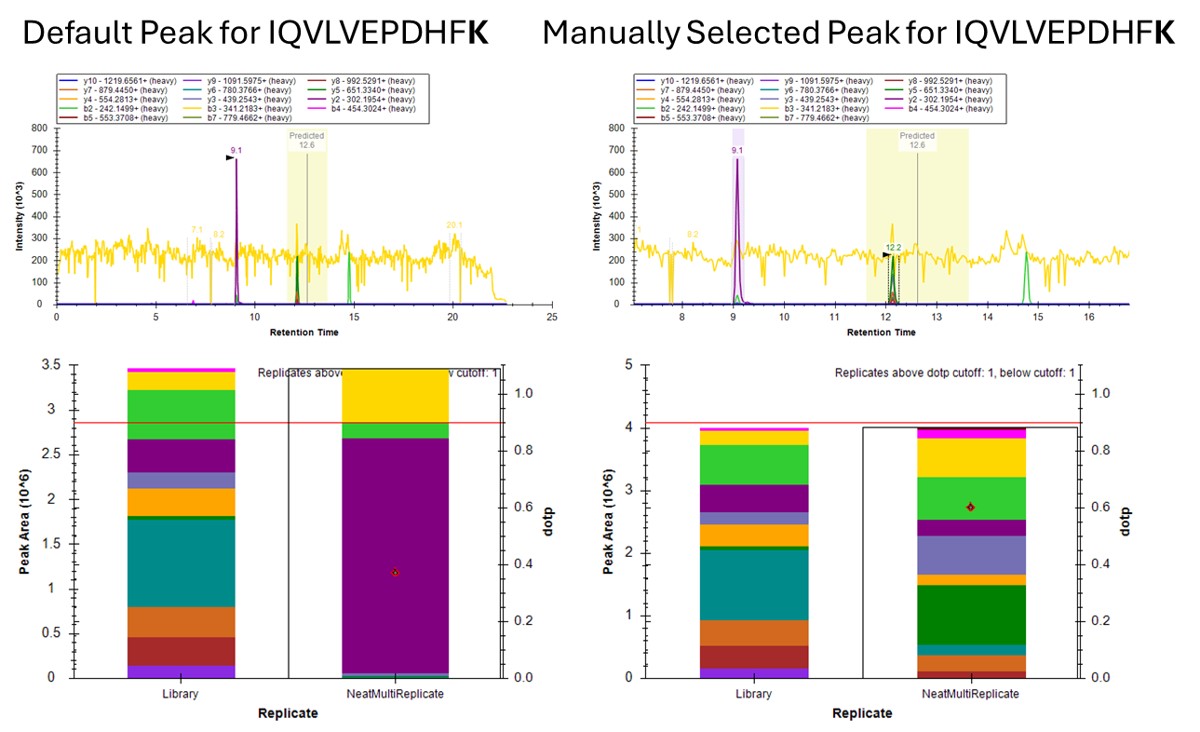
- The Residual Score-to-Run plot is very useful for this step to flag any potential missed peaks. Here we want to click on any of the pink "outlier" dots and inspect them. For example the IQVLVEPDHFK was picked at 9.1 minutes, but the peak in the predicted window at 12.6 has a better library dot product match. Also based on the 100 SPD data, we think that the LLQDSVDFSLADAINTEFK is probably the low intensity peak at 21.6 minutes and not the larger peak at 9.3 minutes that is far away from the predicted RT.
Using Dot Products
-
Another way to find picked peak issues like this is to view a Document grid report that has the dot product scores. Use View / Document Grid (or Alt + 3) to bring up the Document grid, and dock it in the same window as the Retention Time Score-to-Run.
- If you ever can't dock a window where you want it, dock the window in some place that Skyline lets you, and then you will be able to dock it where you originally wanted.
-
Click the Precursor Report, then Customize Report to bring up a dialog menu. Erase columns from the right hand side and then click the binoculars and type 'Dot', and press Find Next until you find Library Dot Product. Select this column to add it to the right hand side, and then press okay.
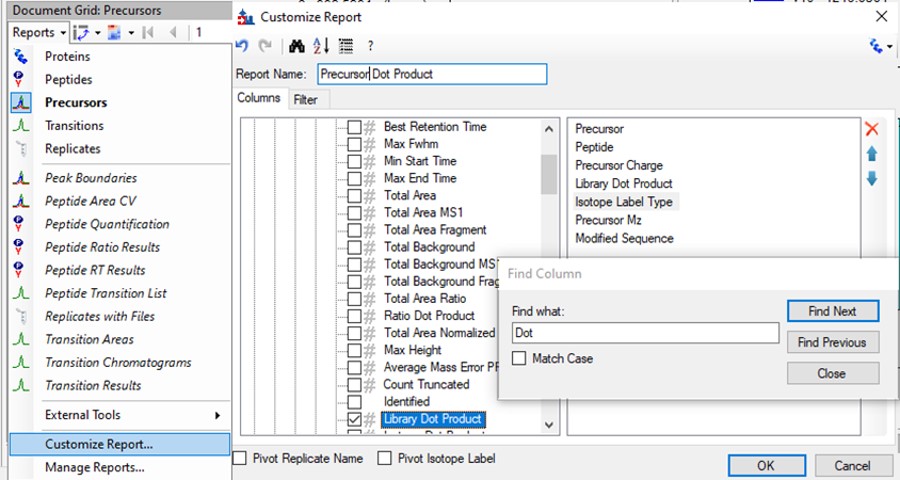
- Now right click on the Library Dot Product column and select Sort Ascending. You can click through on the worse dot product scores to inspect them. Even the ones with poor dot product scores are actually pretty ambiguously belonging to a single precursor at the correct retention time.
Using Peak Picking Models
A final way that can be useful for inspecting this kind of result is to compare the results from the two Skyline peak picking models available at this time. Although in the present case there is not much use for this technique, we'll demonstrate it now.
- Select Refine / Reintegrate. In the Peak scoring model dropdown box, select Add. In the dialog that shows up, keep mProphet as the model, and select Training / Use second best peaks. Using decoys is the method used when you have DIA data and have added Decoy peptides to the document. For targeted methods, the second best peak for the peptide serves as the null distribution for training the peak model. Click Train Model. In the feature scores there are several scores that are not available because there are no light peptides in the document. Scroll down the see the enabled features. Several features are red because they decreased the classification accuracy. Unselect those features and press Train Model again, and click Okay to exit the Edit Peak Scoring Model dialog.

- In the Reintegrate dialog, Add another peak scoring model. This time select the Default model and press Train Model. Then press Okay, and press Okay again in the Reintegrate dialog. This should give us the original peaks selected when the data were imported.

- Go to Refine / Compare Peak Scoring. Use the Add button to add the two peak models that we just made. Go to the Score Comparison tab, select the Default model for the Compare dropdown and the mProphet model for the "with" dropdown. Click the Show conficts only button. Of the 8 conflictin peaks, the greatest ambiguity is probably the first one, LLGNVLVCVLAR. Here there is a very nice peptide peak at 20.9 minutes, but the one at 20.5 although it is further from the predicted RT, has a better dot product. We selected the 20.9 peak, but potentially this is misassigned. We made the same assignment in the 100 SPD method.

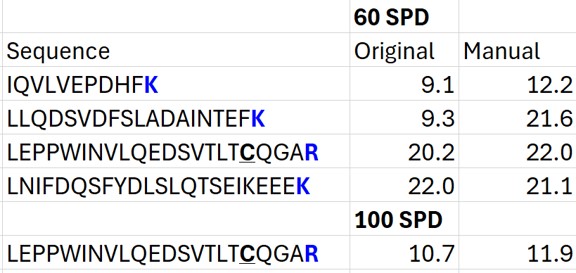
Summary of Changes made to Default Skyline Peak Picking
The figure below summarizes the manual changes we made to the Skyline default peak picking. Note that at the time that we first did the study, Skyline could connect to the Prosit server for library generation. By the time this tutorial was written, Skyline was using something called Koina to do the in silico predictions. While the Prosit models are in theory supported, there was an issue with using them. Therefore there could be some small differences. Note that the most conservative approach would be to search the unscheduled PRM data against a PQ500 .fasta file with static R and C heavy modifications, and only select those peptides that passed some threshold FDR value.
Saving a New, Empirical iRT Calculator
- Now we will update the iRT library so that future experiments can benefit from this curated peak picking. Use Settings / Peptide Settings / Prediction, and click the calculator button to Edit Current. In the dialog that opens, click Add in the bottom right corner and add results.., and an Add iRT Peptides dialog announces that 804 peptide iRT values will be replaced. Click Okay. Click Yes when asked if the iRT Standard values should be recalibrated. Now in the iRT database line, use the name pq500_60spd.irtdb, and give it a new name, PQ500+PRTC 60 SPD, press Okay to exit.

- We've created a new iRT calculator, but have to set it as the active one. Click the calculator icon and Edit List. Press the Add button and then click the dropdown menu, and the new calculator will be there, select that one. Set the Time window to 2 minutes, then press Okay. Select the PQ500+PRTC 60 SPD calculator in the Peptide Settings / Prediction dialog and press Okay to exit to main Skyline window.
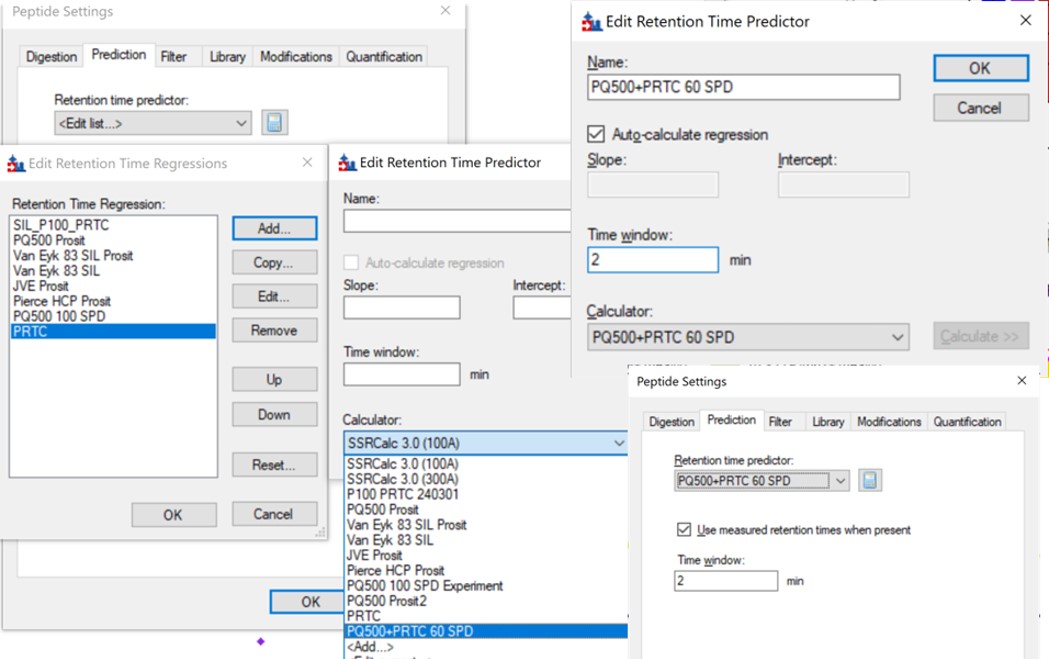
- In the score-to-run plot, right click, select Calculator, and choose the PQ500+PRTC 60 SPD. The plot will update to show an interesting pattern where the errors are all very close to zero. We have now created a new calculator that will make it easier for Skyline to pick peaks next time. We did the exact same set of iRT steps for the 100 SPD Skyline file, because using the new 60 SPD calculator for the 100 SPD does not work perfectly. In the 100 SPD we think that probably the LEPPWINVLQEDSVTLTCQGAR is the peak at 11.9 minutes and not the one at 10.7 minutes.
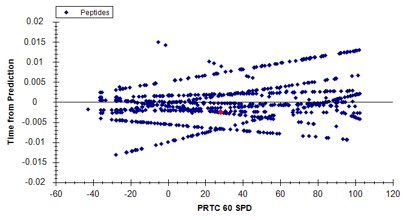
Filtering Transitions with PRM Conductor
This was a neat sample, but we can filter out the transitions that we don't need at this point, with the understanding that when we spike into plasma we may have to refine the transitions even further.
- Select Tools / Thermo / PRM Conductor. The tool opens up. We used the settings in the figure below, where importantly you set Min Good Trans to 5, press Enter, and also select the Keep All Precs box. Press the Send to Skyline button on the bottom left. Save the Skyline document and close the PRM Conductor.

- The number of transitions displayed in the bottom right hand of Skyline will have updated. With the settings we used, the number of transitions dropped from 11,088 to 7,564.

- Use File / Export / Spectral Library and save as Step 2. Neat Unscheduled Multireplicates/PQ500_Neat_60SPD_Refined. Use Settings / Peptide Settings / Library and select Edit List, then Add, and Browse for the file you just saved. Give the library the name PQ500 Neat 60 SPD. Press Okay, and in the Library tab, deselect the PQ500 Koina library and select the new library. Press Okay and save the Skyline document. Make a spectral library in the same way for the 100 SPD document. Close the PRM Conductor instances. We've reached the end of the second step, and are ready to acquire data in plasma.
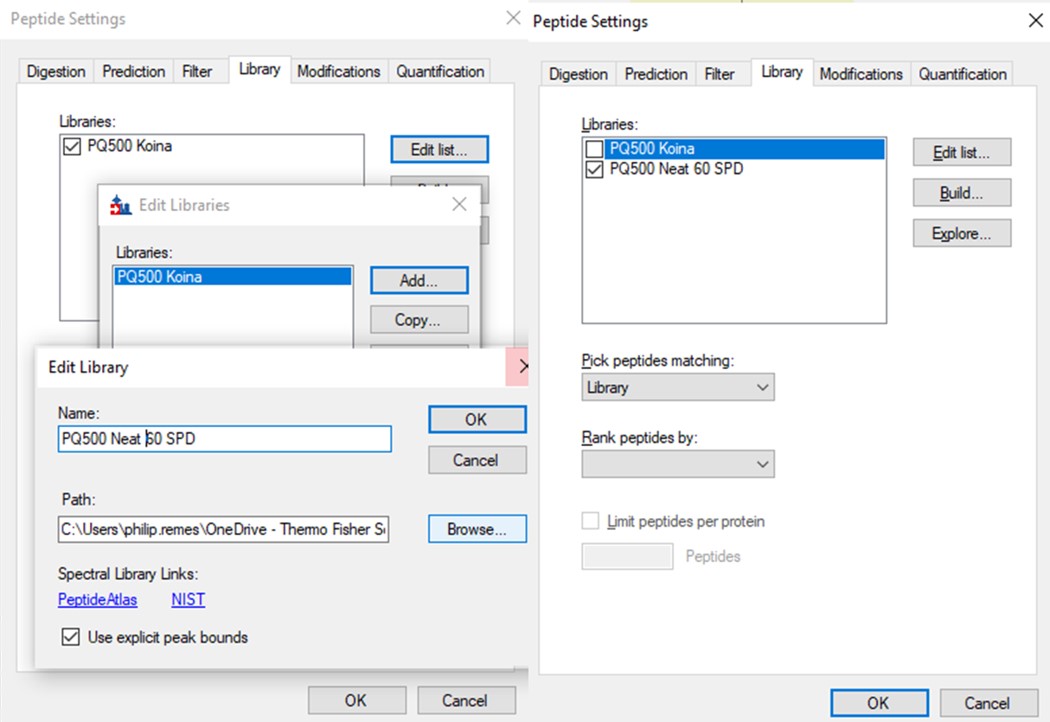
Plasma Wide Window Scheduled
Scheduled PRM for Heavy PQ500 in Plasma with Wide Acquisition Windows
In this step we will create a wide-window PRM method to verify the RT locations of the PQ500 heavy peptides in plasma. Sometimes it can be the case that the RT’s of peptides will be much different when spiked into matrix compared to when analyzed neat. This is expected and likely due to the binding properties of the chromatography stationary phase, which depend on the concentration of analytes in the liquid phase in an equilibrium sometimes referred to as an isotherm. At the end of this section we will have a candidate final method that includes both heavy and light peptides, and that also includes Adaptive RT real-time chromatogram alignment.
Creating Wide Window Methods that Include Adaptive RT Acquisitions
-
Use File / Save As on our files from the last step, pq500_60spd_neat_multireplicate_results_refined.sky and pq500_100spd_neat_multireplicate_results_refined.sky and save in the folder Step 3. Plasma Heavy-Only Wide Window as pq500_60spd_plasma_multireplicate_results.sky and pq500_100spd_neat_multireplicate_results_refined.sky.
-
Use Tools / Thermo / PRM Conductor.
-
Update the settings as in the figure below. After changing any number value, be sure to press the Enter key on the keyboard. The prtc_priority.prot file is selected by double clicking the Protein Priority File text box. This is just a text file with the line “Pierce standards”, the protein name that Skyline gave to the iRT standards. The peptides from any proteins listed in this file (with Skyline's protein names, not accession numbers) are included in the assay, whether or not their transitions meet the requirements. If the Balance Load checkbox is not selected and there are multiple assays to export, each assay will contain the prioritized proteins, and Skyline will be able to use the iRT calculator for more robust peak picking.
-
Note that with the 1.8 minute acquisition window, the right-most plot in PRM Conductor tells us that the 818 precursors in the assay require up to almost 2500 milliseconds to be acquired, and as we have the Balance load box unchecked, they will be split into 2 assays. If Balance Load was checked, then we would create a single assay, for only the precursors that can be acquired in less than the Cycle Time.

-
Enter a suitable Base Name like PQ500_60SPD_Plasma_ToAlign. This reflects the fact that we are including acquisitions to perform Adaptive RT, but we are not actually adjusting our scheduling windows in real time. Our neat standards were not suitable for performing aligning in the complex plasma matrix background.
-
Double click the Method Template field and select the Step 3. Plasma Heavy-Only Wide Window/PQ500_60SPD_ToAlignTemplate.meth file. This file is standard targeted method for Stellar, with 3 experiments. The first is the Adaptive RT DIA experiment, which is being used to gather data for real-time alignment in future targeted methods. The second is a MS1 experiment, which isn't strictly needed, but enables the TIC Normalization feature in Skyline to be used, and can be helpful for diagnostic purposes. Removing it would save on computer disk space. The tMSn experiment can be simply the default tMSn experiment, where we ensure that Dynamic Time Scheduling is Off. If it were on, then PRM Conductor would try to embed alignment spectra from the current data set into the method. Here we leave it off.

-
Press the Export Button. PRM Conductor will open a progress bar and do some work to export a .sky file, and two .meth files with the names PQ500_60SPD_Plasma_ToAlign_0.sky and PQ500_60SPD_Plasma_ToAlign_1.sky. The new .sky file has our new transition list imported and sets the Acquisition mode to PRM. This can be useful especially when discovery data is acquired in a DIA mode, however in this case we want to still compare our neat PQ500 data with the spiked plasma data we’ll be collecting, so we’ll continue using our file pq500_60spd_plasma_multireplicate_results.sky.
-
Do the same thing for the 100 SPD method. Here we can create 3 methods if the LC Peak Width is set to 8. Change the Base Name and Method Template to the 100 SPD versions and press Export Files.
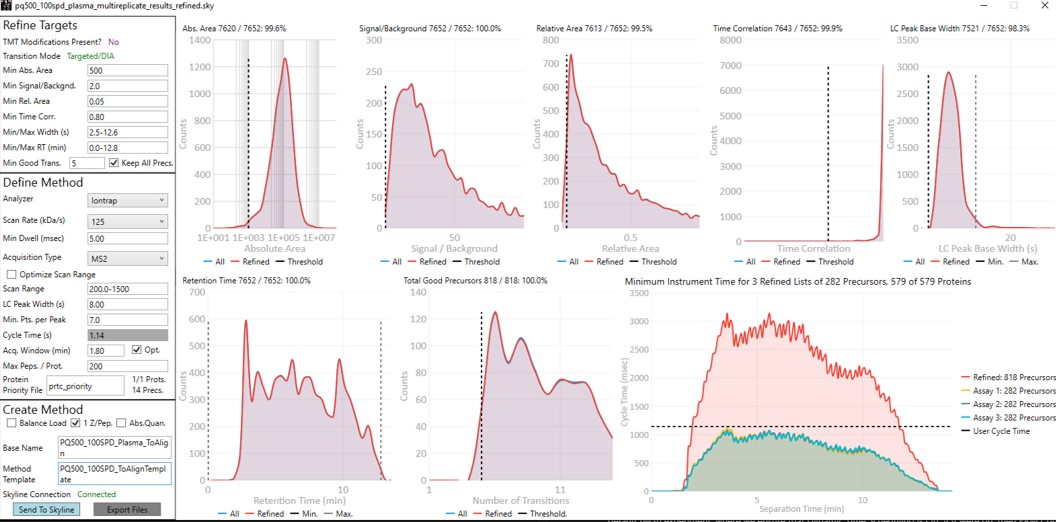
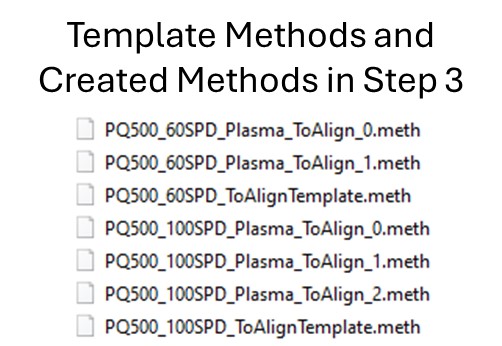
- The created methods in this step are as in the figure below. We are now ready to acquire data using these methods. We did this by spiking in PQ500 at the Biognosys recommended concentration into 300 ng of digested plasma that we got from Pierce. Pierce will offer this as a product for sale in the near future (this is written mid-2024).

Reviewing the Wide Window, Plasma, Scheduled PRM Results
-
Open Settings / Transition Settings and set the Retention time filtering option to Use only scans within 1 minutes of MS/MS IDs. You want to be careful with this filtering because if the RT shifts were greater than +/- 1 minute, some data could be missing. You can always use one number and then change it, and use Edit/Manage Results, and select the replicate and Reimport, to use a wider or narrower filter. In this case the IDs are coming from the spectral library that we created in the previous step. Alternatively one could use Use only scans with X minutes of predicted RT option. We need some kind of RT filtering of this sort to help Skyline differentiate between the iRT peptide SSAAPPPPPR and the PQ500 peptide FQASVATPR, which have the same exact m/z.
-
We acquired data for PQ500 spiked into 300 ng of plasma and put the resulting .raw files in the folder Step 3. Plasma Heavy-Only Wide Window\Raw. Load the results with File / Import Results / Add one replicate, use a name like PlasmaMultireplicate, and select the .raw files for the appropriate 60 or 100 SPD throughput. Skyline will load these data files as a single replicate.
-
Use View / Arrange Graphs / Row so that we can view the Neat and the Plasma replicates at the same time. Right click a chromatogram plot and use Auto Zoom X Axis / None, so that we are zoomed out as far as we can go.
-
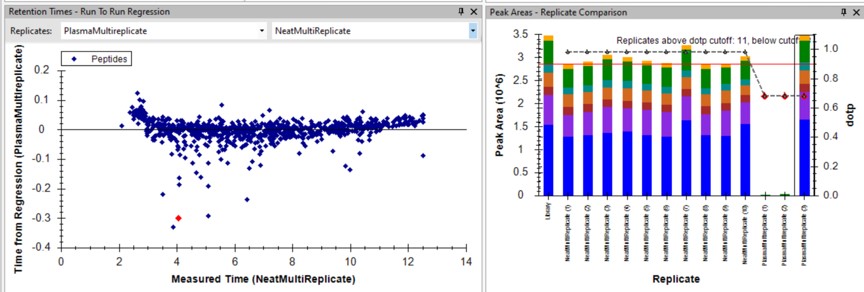
Use View / Retention Times / Regression / Run-to-Run. Your Skyline document should look something like the figure below.
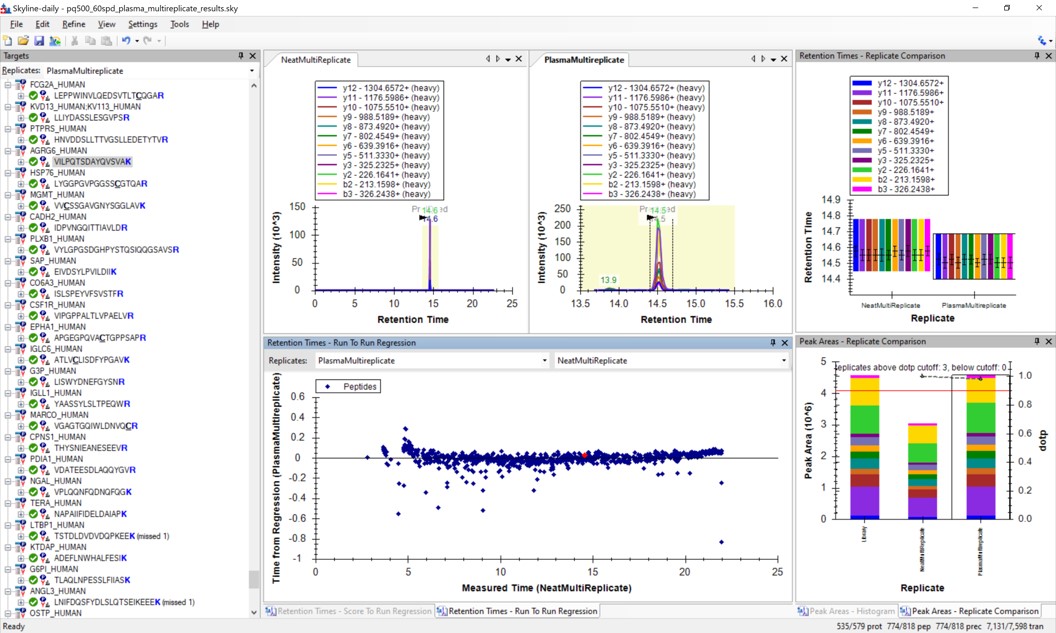
-
Use Save As to save a new version of this file in case we make any changes to the picked peaks. You can use pq500_60spd_plasma_multireplicate_results_refined.sky.
-
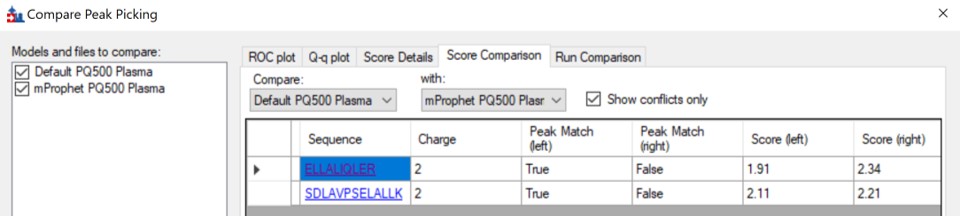
We can do the same steps as above. Make an mProphet and Default Peak picking models with Refine / Reintegrate. Then use Refine / Compare Peak Scoring, select the two new models with the Add button, then select the Score Comparison tab. Select the two models, and click Show conflicts only. We see two descrepancies, and changed the ELLALIQLER peptide from 20.0 to 20.2 minutes, which had a higher dot product and better predicted RT. We kept the default peak for the other peptide.
-
Click on the various outliers and make sure that the peak area plots are showing a good correspondance of the transitions with a high dot product.
-
Use a report with the Library Dot Product sorted from Low-to-High and investigate the worst cases. Even the lowest dot product cases look okay to us.
-
For the 100 SPD data, investigating the Plasma-to-Neat retention times has a similar patterns as for the 60 SPD. There is one case, the FQASVATPR peptide, that has the same exact m/z as the SSAAPPPPPR iRT peptide, which elutes at a similar RT. Reducing the Settings / Transition Settings / RT filtering time to +/- 0.5 minutes can separate these two peptides. We kept all the Skyline picked LC peaks for the 100 SPD document, and saved a new file pq500_100spd_plasma_multireplicate_results_refined.sky.
Creating a Final Method
-
Remove the NeatMultiReplicate using Edit / Manage Results. It's easy to forget to do this. We don’t want PRM Conductor to consider the neat peaks, which are already very clean. Save the Skyline file again.
-
Launch PRM Conductor to clean up interferences and create a final method. Set Min. Good Trans. 5 and check Keep All Precs. Set Min Dwell 5 msec. Set LC Peak Width 11, Min. Pts. Per Peak 7, Acquisition Window 0.6 minutes, and check the Opt. box. This option increases the acquisition windows slightly, especially at the start of the experiment, without going over the user's Cycle Time. Select the prtc_priority.prot file, which in in this case just makes sure that those peptides can't get filtered. Check the Balance Load, 1 Z/prec., and Abs. Quan boxes. This last option instructs the Export command to include light targets for each of the heavy targets. Set a Base Name PQ500_60SPD_Align, and select the PQ500_60SPD_AlignTemplate.meth.
-
This template method is the same as the ToAlign version, only the Dynamic Time Scheduling is set to Adaptive RT. Now when PRM Conductor exports a method, it will compress the qualifying alignment acquisitions in the data and embed them into the created method.
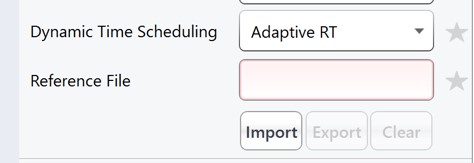
-
We have a small issue here in that there more refined targets (red trace) than we can target. We have to trick PRM Conductor here and set the LC Peak Width to 20 so that all targets are exported, then in the created file change the LC Peak width back to 11 and points per peak to 7. In the future we'll allow the user to just export an "invalid" method.
-
Press Export Files to create the new instrument method.

- Open the PQ500_60SPD_Align.meth file, change the LC Peak width from 20 to 11, and the Points per Peak to 7. Note that the Adaptive RT Reference file has a file name that starts with Embedded, and the Mass List table has entries with heavy peptides having names ending in [+10] or [+8] followed by the corresponding light peptides.

-
Use the Send to Skyline to filter the remaining few poor transitions from these targets, and save the Skyline document state. Export a spectral library like we did before, giving it a name like PQ500_60SPD_Plasma. Configure this library in Settings / Peptide Settings / Library. This is the end of step 3.
-
For the 100 SPD case, from the pq500_100spd_plasma_multireplicate_results_refined.sky file, Launch PRM Conductor. Check the Optimize Scan Range box. This will produce targets with customized scan ranges for each target, significantly increasing the acquisition speed, at the cost of some injection time and sensitivity. Set an appropriate Base Name and select the PQ500_100SPD_AlignTemplate.meth for the Method Template. Use the same trick as for the 60 SPD, setting the LC peak width to 20 seconds and Export the method. Then open the method that is created and change the LC peak width back to 7 with 6 points per peak.
-
Press the Send to Skyline button. Export the spectral library and configure it in the Peptide Settings / Library tab. Save the pq500_100spd_plasma_multireplicate_results_refined.sky.

- We now have candidate final methods created for 60 SPD and 100 SPD, respectively PQ500_60SPD_Align.meth and PQ500_100SPD_Align.meth. We are ready to move to the next step and validate the assay.
Plasma Narrow Windows Scheduled
Scheduled PRM for Light-Heavy PQ500 in Plasma with Narrow Acquisition Windows
Analysis of the Heavy Peptides
In step 3 we created two candidate final methods for the 60 and 100 SPD assays. Take the file pq500_60spd_plasma_multireplicate_results_refined.sky and pq500_1000spd_plasma_multireplicate_results_refined.sky, and resave them in the folder Step 4. Plasma Light-Heavy Narrow Window, with names like pq500_60spd_plasma_final_replicates.sky and _pq500_100spd_plasma_final_replicates.sky. In this step we’ll analyze the results of the light/heavy methods created in Step 3. Of particular interest will be the histogram of coefficient of variance values for the peak areas.
-
Use File / Import Results / Add single-injection replicates in files and press Okay. Select the 10 files in Step 4. Plasma Light-Heavy Narrow Window\Raw\60SPDReplicates and press Open. Remove the common prefix and press Okay to load the results. Remove the PlasmaMultiReplicate with Edit / Manage Results and Save the document.
-
Select View / Peak Areas / CV Histogram. The CV histograms have ~94% of the targets with CV < 20%, with medians of 3.8 and 4.9%, which are excellent.

You can click on the histogram, which will open a Find Results window with some of the peptides that are close in CV to the value that was pressed. Double clicking any peptide sequence in the Find Results table will make that peptide active, whereupon one can check the peak shape, peak area, and retention time variations for the 8 replicates. Many/most peptides have results like LFGPDLK below.
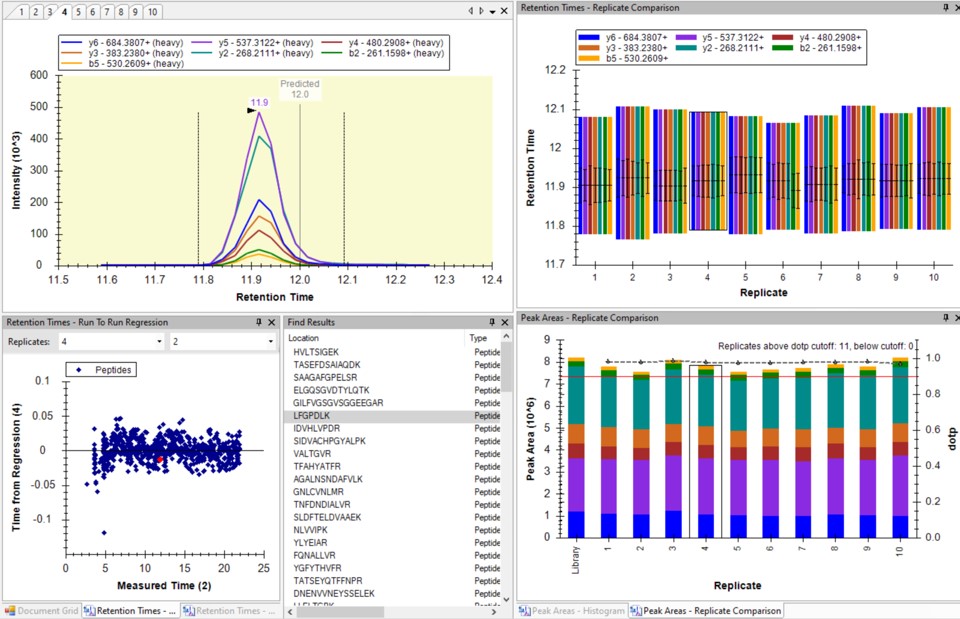
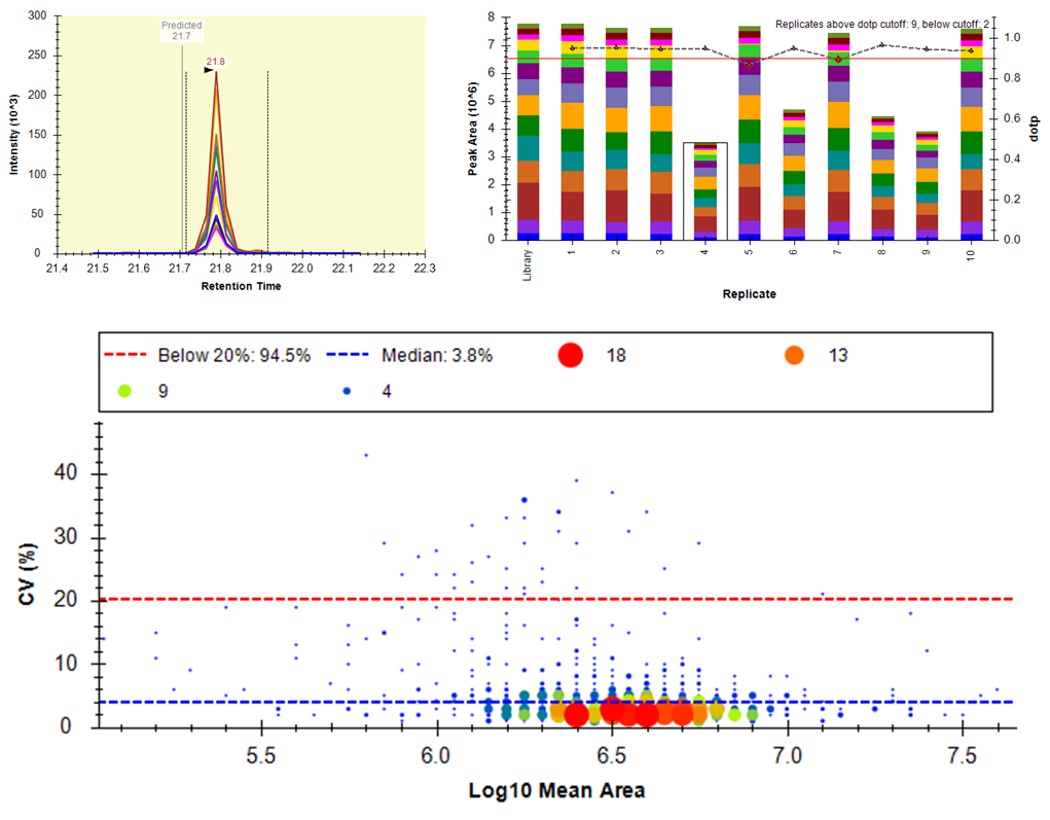
- Another interesting plot is the 2D CV Histogram, found under the View / Peak Areas menu. Here we see that there are 45 targets with CV > 20%. Clicking on the dots makes the peptide in question active. An example peptide SLADELALVDVLEDK is shown below, that has > 20% CV. It has a very skinny peak shape, eluting during the column wash portion of the run. Our analyses show that peptides very early and late in the assays have a high probability of have poor CV. One way to filter these peptides out is based on their narrow peak shape. PRM Conductor can set a minimum LC peak width bounds to do this.
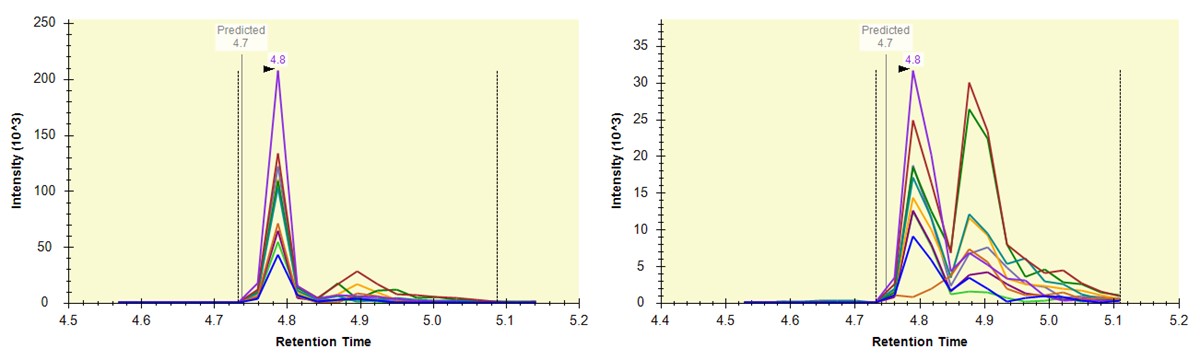
- Some peptides have bad CV's because they have interference an interference that varies from one replicate to another, like ANHEEVLAAGK below.
- One could simply filter all the peptides with a CV greater than a threshold from the document, using Refine / Advanced / Consistency, and set 20% in the CV cutoff box. Or one could try using MS3 acquisition in PRM Conductor, to save those peptides with poor CV's.
Analysis of Both Light and Heavy Peptides
-
Now we will add in the light precursors, that were measured but currently are not in the Skyline document. Save the document and then Save again with the names pq500_60spd_plasma_final_lightheavy_replicates.sky and pq500_100spd_plasma_final_lightheavy_replicates.sky.
-
Use Refine / Advanced, and select the Add box. The Remove label type combo box title changes to Add label type. Select light and press Okay to close the Refine dialog. Each peptide will now have its light precursor added. Use Edit / Manage Results, select all the replicates and press the Reimport button.
- Use View / Peak Areas / 2D CV Histogram, and set Normalized to Heavy. The PQ500 standards were designed to look for a set of proteins of interest, not all of which are expressed in normal plasma. For this reason many of the Light / Heavy area ratios are low, and have high coefficient of variance.

- Some peptides, like AGALNSNDAFVLK have significant endogenous peptide, and therefore have very good CV. Other peptides like EILVGDVGQTVDDPYATFVK have no observable endogenous peptide, and the area ratio therefore has a much higher CV. This general trend is observed in the CV histogram above.

-
This is the end of Step 4. We've demonstrated how to analyze replicate data for absolute quantitation with light and heavy peptides. A next step that some users will want to perform is a dilution curve. For absolute quantitation this takes two forms.
- Heavy Dilution: Constant endogenous (light) peptide amount, varying heavy spike-in concentration
- Light Dilution: Constant heavy spike-in concentration, varying endogenous sample.
-
The Light Dilution is a little easier to perform, because with the Settings / Peptide Settings / Modifications / Internal standard type is set to heavy, and thus Skyline uses the integration boundaries of the heavy peptides to integrate the light signals and determine whether the light/heavy ratios are sufficient for quantitation.
-
The Heavy Dilution is difficult, because eventually Skyline can't find the heavy peptide signal, and doesn't keep a constant integration boundary. Sometimes Skyline will jump over to the next biggest LC peak and ruin the dilution curve. We have sometimes used a script to set constant integration boundaries and solve this issue.
-
Calculating LOQs and LODs for large scale assays is still a little difficult, and we have used python scripts to do this. Skyline is also working on making improvements, and there will be updates in the future. We are submitting a paper soon that will have links to these scripts, for the intrepid that might be interested in exploring them.